Engineering Isoprenoid Biosynthesis: Pathways, Progress, and Pharmaceutical Applications
This article provides a comprehensive overview of the latest advances in isoprenoid biosynthetic pathway engineering, a field critical for the sustainable production of valuable therapeutics, nutraceuticals, and biofuels.
Engineering Isoprenoid Biosynthesis: Pathways, Progress, and Pharmaceutical Applications
Abstract
This article provides a comprehensive overview of the latest advances in isoprenoid biosynthetic pathway engineering, a field critical for the sustainable production of valuable therapeutics, nutraceuticals, and biofuels. Tailored for researchers, scientists, and drug development professionals, it explores the foundational biology of the mevalonate (MVA) and methylerythritol phosphate (MEP) pathways, details cutting-edge metabolic engineering methodologies in both model and non-model organisms, addresses common optimization challenges and troubleshooting strategies, and offers a comparative analysis of microbial production platforms. The synthesis of these perspectives aims to guide future engineering efforts toward more efficient and commercially viable biomanufacturing of complex isoprenoid compounds.
Deconstructing the Core: Foundational Pathways in Isoprenoid Biosynthesis
Isoprenoids, also known as terpenoids, represent the most diverse class of natural products in nature, with essential biological functions including hormone signaling, antioxidation, electron transport, and cell wall biosynthesis [1]. All isoprenoids are synthesized from two universal five-carbon (C5) precursors: isopentenyl diphosphate (IPP) and its isomer dimethylallyl diphosphate (DMAPP) [1] [2]. The biosynthetic routes to these central metabolites proceed via two distinct, evolutionarily conserved pathways: the mevalonate (MVA) pathway and the methylerythritol phosphate (MEP) pathway [1] [2].
The MVA pathway was the first to be discovered and was long considered the sole route to isoprenoids [1]. The MEP pathway was identified more recently, resolving long-standing biochemical inconsistencies in certain bacteria and plants [1]. These pathways differ in their reaction mechanisms, evolutionary distribution, cellular localization, and susceptibility to regulation and inhibition. This Application Note provides a comprehensive overview of both pathways, detailing their biochemical reactions, experimental protocols for investigation, and their application in metabolic engineering for the production of high-value isoprenoids.
Pathway Biochemistry and Distribution
The Mevalonate (MVA) Pathway
The canonical MVA pathway, predominantly found in the cytoplasm of eukaryotes, archaea, and some gram-positive bacteria, initiates from acetyl-CoA [3] [2]. It involves a series of six enzymatic steps to produce IPP:
- Condensation: Two acetyl-CoA molecules condense to form acetoacetyl-CoA, catalyzed by acetoacetyl-CoA thiolase (AACT).
- Second Condensation: Acetoacetyl-CoA combines with a third acetyl-CoA to form 3-hydroxy-3-methylglutaryl-CoA (HMG-CoA), a reaction catalyzed by HMG-CoA synthase (HMGS).
- Reduction: HMG-CoA is reduced to mevalonate by HMG-CoA reductase (HMGR). This is the first committed and rate-limiting step of the pathway [4].
- Phosphorylation: Mevalonate is sequentially phosphorylated by mevalonate kinase (MVK) and phosphomevalonate kinase (PMK) to yield mevalonate-5-diphosphate.
- Decarboxylation: Mevalonate-5-diphosphate is decarboxylated to form IPP, catalyzed by diphosphomevalonate decarboxylase (DMD/MVD) [2].
- Isomerization: IPP is isomerized to DMAPP by IPP isomerase (IDI) [2].
A modified, archaeal MVA pathway has been discovered that utilizes different enzymes—phosphomevalonate dehydratase (PMDh), anhydromevalonate phosphate decarboxylase (AMPD), and isopentenyl phosphate kinase (IPK)—for the final steps from mevalonate-5-phosphate to IPP. This modified pathway consumes one less ATP molecule than the canonical pathway, offering a potential energy-saving advantage for metabolic engineering [5].
The Methylerythritol Phosphate (MEP) Pathway
The MEP pathway is utilized by most prokaryotic microorganisms, the plastids of plants, and some lower eukaryotes like apicomplexan parasites [1] [6]. It starts from the precursors pyruvate and D-glyceraldehyde 3-phosphate (GAP) and proceeds through seven enzymatic steps:
- Condensation: Pyruvate and GAP are condensed to form 1-deoxy-D-xylulose 5-phosphate (DXP), catalyzed by DXP synthase (DXS) [1]. DXP also serves as a precursor for thiamine and pyridoxal biosynthesis [1].
- Reduction and Isomerization: DXP is converted to 2-C-methyl-D-erythritol 4-phosphate (MEP) by DXP reductoisomerase (DXR). This is the first committed step dedicated to isoprenoid biosynthesis in this pathway and a key regulatory point [1] [4].
- Cytidylation: MEP is converted to 4-(cytidine 5'-diphospho)-2-C-methyl-D-erythritol (CDP-ME) by MEP cytidylyltransferase (IspD).
- Phosphorylation: CDP-ME is phosphorylated to form CDP-ME-2-phosphate (CDP-MEP) by CDP-ME kinase (IspE).
- Cyclization: CDP-MEP is converted to 2-C-methyl-D-erythritol 2,4-cyclodiphosphate (MEcDP) by MEcDP synthase (IspF) [6].
- Reductive Ring Opening: MEcDP is converted to 1-hydroxy-2-methyl-2-(E)-butenyl 4-diphosphate (HMBPP) by HMBPP synthase (IspG). This enzyme, along with IspH, contains oxygen-sensitive iron-sulfur (Fe-S) clusters, making this pathway sensitive to oxidative stress [6].
- Final Reduction: HMBPP is reduced to a mixture of IPP and DMAPP in a ~5:1 ratio by HMBPP reductase (IspH) [2]. Unlike the MVA pathway, DMAPP is synthesized directly and does not solely rely on the isomerization of IPP.
A unique aspect of the MEP pathway is its role as an oxidative stress sensor and response system. The Fe-S cluster enzymes IspG and IspH are sensitive to reactive oxygen species (ROS). Under oxidative stress, the pathway intermediate MEcDP accumulates and can act as a stress signaling molecule, potentially linking isoprenoid metabolism to the cell's stress response [6].
Table 1: Comparative Overview of the MVA and MEP Pathways
| Feature | Mevalonate (MVA) Pathway | Methylerythritol Phosphate (MEP) Pathway |
|---|---|---|
| Initial Precursors | Acetyl-CoA (3 molecules) [2] | Pyruvate + Glyceraldehyde-3-phosphate (GAP) [2] |
| Key Intermediates | HMG-CoA, Mevalonate [2] | 1-deoxy-D-xylulose 5-phosphate (DXP), MEP [1] |
| First Committed Step | HMG-CoA to Mevalonate (by HMGR) [1] | DXP to MEP (by DXR) [1] |
| Oxygen Sensitivity | Generally oxygen-independent (eukaryotic version) | Oxygen-sensitive (due to Fe-S cluster enzymes IspG/H) [6] |
| Evolutionary Distribution | Eukaryotes (cytosol), Archaea, some Bacteria [3] [2] | Most Bacteria, Plant Plastids, Apicomplexan parasites [3] [6] |
| Theoretical Yield on Glucose | 25.2% [6] | 30.2% [6] |
| ATP Consumption (per IPP) | 3 ATP (canonical pathway) [4] | Varies; generally lower than MVA |
| Pathway-Specific Inhibitors | Lovastatin, Mevinolin (target HMGR) [1] [7] | Fosmidomycin (targets DXR) [7], Ketoclomazone (targets DXS) [1] |
The following diagram illustrates the sequential steps of both the MVA and MEP pathways, highlighting their subcellular localization and key metabolites.
Experimental Protocols for Pathway Analysis
Protocol: Investigating Pathway Contribution with Stable Isotope Labeling
This protocol is adapted from studies investigating the contribution of the MVA and MEP pathways to specific isoprenoids in complex systems like plants [7].
Purpose: To determine the relative contribution of the MVA and MEP pathways to the biosynthesis of a target isoprenoid (e.g., dolichols, polyprenols, sterols, carotenoids).
Principle: Organisms or cells are fed with stable isotope-labeled precursors that are specific to each pathway. The incorporation of the label into the target isoprenoid is then quantified using Mass Spectrometry (MS) to determine the fractional contribution from each pathway.
Materials:
- Deuteriated MEP Pathway Precursor: (5,5-2H2)-1-Deoxy-D-xylulose (D-DX) [7].
- Deuteriated MVA Pathway Precursor: ((6,6,6-2H3)Methyl)mevalonolactone (D-MVL) [7].
- Internal Standards: Appropriate internal standards for the target isoprenoid (e.g., naphthalene for terpenes, 5-α-cholestan-3-α-ol for sterols) [8].
- Extraction Solvents: Methyl tert-butyl ether (MTBE), chloroform:methanol (2:1 v/v), hexane, petroleum ether.
- Instrumentation: Gas Chromatography-Mass Spectrometry (GC-MS) or Liquid Chromatography-Mass Spectrometry (LC-MS).
Procedure:
- Treatment: Divide the biological material (e.g., plant tissue, cell culture) into experimental groups.
- Group 1 (MEP): Administer deuteriated D-DX.
- Group 2 (MVA): Administer deuteriated D-MVL.
- Group 3 (Control): No treatment or natural abundance precursor.
- Incubation: Incubate the material under defined growth conditions (temperature, light, humidity) for a predetermined period to allow for metabolic incorporation.
- Harvesting and Extraction:
- Flash-freeze the material in liquid nitrogen and homogenize to a fine powder.
- Extract isoprenoids based on their chemical nature.
- For volatile terpenes (e.g., mono/sesquiterpenes): Extract with MTBE spiked with an internal standard [8].
- For sterols: Extract with chloroform:methanol, followed by saponification with KOH/methanol, and partition with hexane [8].
- For carotenoids: Extract with acetone/petroleum ether, followed by saponification and purification [8].
- Analysis:
- Derivatize extracts if necessary for GC-MS analysis.
- Analyze samples by GC-MS or LC-MS.
- Monitor the mass shift of the target isoprenoid ions corresponding to the incorporation of deuterium atoms from the labeled precursors.
- Data Analysis:
- Calculate the abundance and distribution of labeled and unlabeled isotopologues.
- Determine the percentage of the target isoprenoid derived from each pathway based on the incorporation of the respective labeled precursor.
Protocol: Functional Complementation Assay for MEP Pathway Enzymes
This protocol is based on the reconstitution of the essential MEP pathway in engineered E. coli to study enzymes from pathogens like Plasmodium falciparum [9].
Purpose: To establish a platform for screening inhibitors or validating the function of MEP pathway enzymes (e.g., IspH, ferredoxin) in a genetically tractable host like E. coli.
Principle: An E. coli mutant strain that lacks a functional MEP pathway (e.g., a flavodoxin and ispH double mutant) is engineered. The strain is rescued by expressing the target enzyme(s) from a heterologous organism, making bacterial growth dependent on the function of the introduced enzyme(s).
Materials:
- Bacterial Strain: E. coli flavodoxin and ispH double mutant [9].
- Expression Plasmids: Constructs expressing the target MEP pathway genes (e.g., ptFd, ptFNR, and IspH from P. falciparum) under inducible promoters [9].
- Growth Media: Minimal medium (e.g., M9) that forces the bacterium to rely on the MEP pathway for isoprenoid production.
- Chemical Inhibitors: (Optional) For inhibitor screening, e.g., Fosmidomycin.
Procedure:
- Strain Transformation: Co-transform the mutant E. coli strain with the plasmid(s) expressing the heterologous MEP pathway genes.
- Complementation Assay:
- Inoculate transformed colonies into liquid minimal medium with the appropriate inducer (e.g., arabinose or IPTG).
- For inhibitor screening, include the compound at various concentrations.
- Incubate cultures with shaking at 37°C.
- Growth Monitoring:
- Monitor bacterial growth by measuring optical density (OD600) over 24-48 hours.
- Functional expression of the heterologous enzymes will restore the MEP pathway, allowing for bacterial proliferation. Non-functional enzymes or effective inhibitors will result in impaired growth.
- Validation:
- Validate the biochemical consequences by analyzing MEP pathway intermediate accumulation using LC-MS, if growth defects are observed [9].
Metabolic Engineering Applications
The two pathways offer distinct advantages and have been extensively engineered in various microbial and plant hosts to enhance the production of valuable isoprenoids, such as lycopene, β-farnesene, and artemisinin [5] [10] [4].
A common strategy involves introducing the entire MVA pathway into native MEP pathway hosts like E. coli or cyanobacteria to create a redundant, high-flux source of IPP and DMAPP [5] [4]. Recent work has focused on optimizing these pathways by using enzymes from different sources to reduce energy consumption (ATP) or bypass regulatory bottlenecks [5].
Table 2: Key Research Reagent Solutions for MVA and MEP Pathway Engineering
| Reagent / Tool | Function / Target | Application in Research |
|---|---|---|
| Fosmidomycin | Potent inhibitor of DXR enzyme in MEP pathway [7] | Probing MEP pathway flux; selecting for engineered strains with alternative pathways [7] |
| Mevinolin (Lovastatin) | Inhibitor of HMGR enzyme in MEP pathway [1] [7] | Probing MVA pathway flux; selecting for engineered strains with enhanced MVA activity [7] [8] |
| Ketoclomazone | Inhibitor of DXS enzyme in MEP pathway [1] | Chemical tool to suppress the MEP pathway in plants and bacteria [1] |
| Deuteriated Precursors (D-DX, D-MVL) | Pathway-specific labeled precursors [7] | Tracing carbon flux through MVA and MEP pathways; quantitative metabolic labeling studies [7] |
| Archaeal MVA Pathway Genes (PMDh, AMPD, IPK) | Energy-saving alternative route from mevalonate to IPP [5] | Metabolic engineering to reduce ATP burden and enhance isoprenoid yields in heterologous hosts [5] |
| 2A Peptide-Linked Polyproteins | Tool for stoichiometric, co-expression of multiple genes (e.g., HMGR and PMK) [8] | Overcoming regulatory bottlenecks in engineered pathways by ensuring balanced enzyme expression [8] |
The following diagram outlines a generalized metabolic engineering workflow for enhancing isoprenoid production, incorporating strategies such as heterologous pathway expression and enzyme engineering.
Concluding Remarks
The MVA and MEP pathways are fundamental to isoprenoid biosynthesis, each with unique biochemical and regulatory characteristics. The choice between engineering, inhibiting, or utilizing these pathways depends on the target organism and the desired application. The MEP pathway offers a higher theoretical carbon yield and is a target for antibacterial and herbicide development, but its oxygen sensitivity presents an engineering challenge [6]. The MVA pathway, particularly its energy-efficient archaeal variants, holds great promise for metabolic engineering in industrial microbes [5]. Future research will continue to elucidate the complex cross-talk between these pathways in plants and develop more sophisticated engineering strategies to harness their full potential for the sustainable production of high-value isoprenoids.
Key Enzymes and Regulatory Nodes in Precursor Synthesis
Isoprenoids represent one of the largest and most diverse classes of natural products, with over 70,000 identified compounds performing essential functions across all domains of life [11]. These metabolites have significant commercial, pharmacological, and agricultural value, with applications spanning pharmaceuticals, nutraceuticals, fragrances, colorants, and advanced biofuels [10] [11]. A fundamental challenge in isoprenoid research involves understanding and engineering the key enzymes and regulatory nodes that control the synthesis of universal C5 isoprenoid precursors, isopentenyl diphosphate (IPP) and dimethylallyl diphosphate (DMAPP) [10] [12]. This application note details the core enzymatic machinery, regulatory mechanisms, and experimental methodologies essential for advanced research and engineering of isoprenoid precursor synthesis, providing a structured framework for scientists engaged in metabolic engineering and drug development.
Isoprenoid Biosynthetic Pathways: Topology and Key Enzymes
All isoprenoids originate from two universal C5 building blocks, IPP and DMAPP. Nature employs two distinct pathways for their synthesis, each with unique enzymatic components, cofactor requirements, and regulatory characteristics [10] [11].
Table 1: Core Pathways for Isoprenoid Precursor Synthesis
| Feature | Methylerythritol Phosphate (MEP) Pathway | Mevalonate (MVA) Pathway | Isopentenol Utilization Pathway (IUP) |
|---|---|---|---|
| Organism Distribution | Most bacteria, plastids of plants and algae [10] [11] | Most eukaryotes, archaea, and certain bacteria [10] [11] | Synthetic pathway [12] |
| Subcellular Localization | Plastids (in plants/algae) [10] | Cytosol (in plants) [10] | Cytosol |
| Initial Substrates | Pyruvate + Glyceraldehyde-3-phosphate (G3P) [10] [12] | Acetyl-CoA [10] [12] | Isoprenol or Prenol [12] |
| Key Intermediate | 2-C-Methyl-D-erythritol-2,4-cyclopyrophosphate (MEcPP) [13] | Mevalonic Acid [12] | Isopentenyl Monophosphate (IP) / Dimethylallyl Monophosphate (DMAP) [12] |
| Number of Enzymatic Steps | 7 [12] | 7 [12] | 2 [12] |
| Primary Cofactor Requirements | ATP, CTP, NADPH [12] | 2 NADPH, 3 ATP [12] | 2 ATP [12] |
| End Products (IPP:DMAPP Ratio) | ~1:5 [12] | IPP only (DMAPP via IdI) [12] | Dependent on alcohol isomer input [12] |
The MEP Pathway
The MEP pathway initiates from the condensation of pyruvate and glyceraldehyde-3-phosphate, catalyzed by DXP synthase (DXS) – a key regulatory node [12]. This pathway features several oxygen-sensitive iron-sulfur cluster enzymes (IspG and IspH) whose inactivation under oxidative stress can lead to intermediate accumulation and carbon loss [12]. The intermediate MEcPP has been identified as a stress signaling molecule in bacteria and plants [13]. Feedback inhibition of DXS by IPP represents a critical regulatory constraint for metabolic engineering [12].
The MVA Pathway
The classical MVA pathway begins with the sequential condensation of two acetyl-CoA molecules, regulated by HMG-CoA synthase and HMG-CoA reductase (HMGR) [12]. HMGR is considered the primary rate-limiting enzyme and is subject to complex feedback inhibition by downstream isoprenoids, including IPP, DMAPP, and farnesyl diphosphate (FPP) [12]. Some bacteria, including the pathogen Mycobacterium marinum, encode both MEP and MVA pathways, with the MVA pathway potentially providing metabolic flexibility under stress conditions [13].
Engineered and Synthetic Pathways
The Isopentenol Utilization Pathway (IUP) is a synthetic, two-step pathway that uses isoprenol or prenol as substrates, requiring only ATP as a cofactor [12]. This pathway decouples isoprenoid synthesis from central carbon metabolism, avoids native regulatory bottlenecks, and can sustain very high flux, making it a valuable tool for metabolic engineering [12]. Key enzymes include promiscuous kinases like choline kinase from Saccharomyces cerevisiae (ScCK) for the first phosphorylation and isopentenyl phosphate kinase from Arabidopsis thaliana (AtIPK) for the second step [12].
Critical Regulatory Nodes in Precursor Synthesis
Fine-tuned regulation ensures proper carbon flux through isoprenoid pathways, balancing primary and secondary metabolic demands.
Table 2: Key Regulatory Nodes and Metabolic Engineering Strategies
| Regulatory Node/Enzyme | Pathway | Regulatory Mechanism | Engineering Strategy |
|---|---|---|---|
| DXP Synthase (DXS) | MEP | Feedback inhibition by IPP [12] | Protein engineering to alleviate feedback inhibition; promoter engineering to enhance expression [10] |
| HMG-CoA Reductase (HMGR) | MVA | Multilevel feedback inhibition by CoA, NAD(P)+, IPP, DMAPP, GPP, FPP [12] | Use of deregulated enzyme variants; compartmentalization to separate from inhibitors [10] |
| IspG/IspH Enzymes | MEP | Oxygen sensitivity via iron-sulfur clusters; pathway intermediate excretion under stress [12] | Cultivation under controlled oxygenation; use of oxygen-stable enzyme orthologs [12] |
| MEP/MVA Pathway Crosstalk | Both | Compensatory changes in intermediate concentrations when one pathway is modulated [13] | Knockout of competing pathways; fine-tuning of dual-pathway expression [10] [13] |
| Choline Kinase (ScCK) in IUP | IUP | Limited catalytic efficiency for prenol (KM=1,113 μM, kcat=1.1 s⁻¹) [12] | Enzyme engineering to improve kinetics; screening of kinase libraries for enhanced activity [12] |
Beyond feedback inhibition, recent research has revealed unexpected regulatory connections. In Arabidopsis, the isoprenoid biosynthesis enzyme 1-hydroxy-2-methyl-2-(E)-butenyl-4-diphosphate synthase (HDS), which catalyzes the penultimate step in the MEP pathway, also participates in chloroplast RNA editing by physically interacting with and modulating the dimerization of RNA editing factors MORF2 and MORF9 [14]. This provides a direct molecular link between isoprenoid metabolism and organellar gene expression.
Experimental Protocols for Analyzing Precursor Synthesis
Protocol for Intracellular Metabolite Analysis in Microbial Systems
This protocol enables quantitative analysis of isoprenoid pathway intermediates in model microorganisms including E. coli, S. cerevisiae, P. tricornutum, C. reinhardtii, and Synechocystis sp. PCC 6803 [11].
Materials and Reagents
- Quenching Solution: 60% methanol in water (v/v), pre-chilled to -40°C
- Extraction Solvent: 75% ethanol with 0.1 M formic acid, pre-heated to 95°C
- LC-MS Solvents: HPLC-grade water, methanol, acetonitrile
- Internal Standards: Stable isotope-labeled analogs of target metabolites (when available)
- Solid Phase Extraction Cartridges: Oasis HLB or similar for metabolite cleanup
Procedure
- Culture and Sampling: Grow microorganisms under standardized conditions. For sampling, rapidly transfer culture aliquots (1-2 mL) into pre-chilled quenching solution (5x volume) to immediately halt metabolism [11].
- Quenching and Washing: Maintain samples at -40°C for 3 minutes, then centrifuge at high speed (14,000 × g, 5 min, -20°C). Remove supernatant and wash cell pellet with cold phosphate-buffered saline [11].
- Metabolite Extraction: Add pre-heated extraction solvent to cell pellet, vortex vigorously, and incubate at 95°C for 5 minutes with occasional mixing. Centrifuge to remove debris and collect supernatant [11].
- Sample Concentration: Dry supernatants under nitrogen or vacuum centrifugation. Reconstitute in appropriate LC-MS compatible solvent for analysis [11].
- LC-MS/MS Analysis:
- For IPP/DMAPP Separation: Use a chiral column (e.g., CHIRALPAK IA-3) with isocratic elution (10 mM ammonium acetate in 70:30 methanol:water) for baseline separation [11].
- For Broader Metabolite Profiling: Use a C18 column with gradient elution (water/methanol with 10 mM ammonium acetate) for partial separation of IPP/DMAPP while capturing other intermediates [11].
- MS Detection: Employ negative ion mode electrospray ionization with multiple reaction monitoring (MRM) for optimal sensitivity [11].
Critical Steps and Troubleshooting
- Maintain consistent processing speed across samples to minimize technical variability
- For microorganisms with robust cell walls (e.g., microalgae), incorporate a bead-beating step during extraction
- Include quality control samples pooled from all experimental conditions
- Monitor for potential isomerization during extraction and analysis
Metabolic Flux Analysis Using Stable Isotope Tracers
Stable isotope labeling enables quantitative analysis of pathway utilization and flux distribution [15].
Procedure
- Labeling Strategy: Introduce ¹³C- or ¹⁵N-labeled precursors (e.g., [U-¹³C]glucose, [1-¹³C]pyruvate, or [¹⁵N]anthranilate) during active growth phase [15].
- Time-Course Sampling: Collect samples at multiple time points following tracer introduction to capture kinetic flux profiles [15].
- Sample Processing: Follow metabolite extraction protocol above.
- Mass Spectrometry Analysis: Use high-resolution LC-MS to resolve isotopomer distributions. Calculate fractional enrichment and apply computational modeling to determine metabolic flux [15].
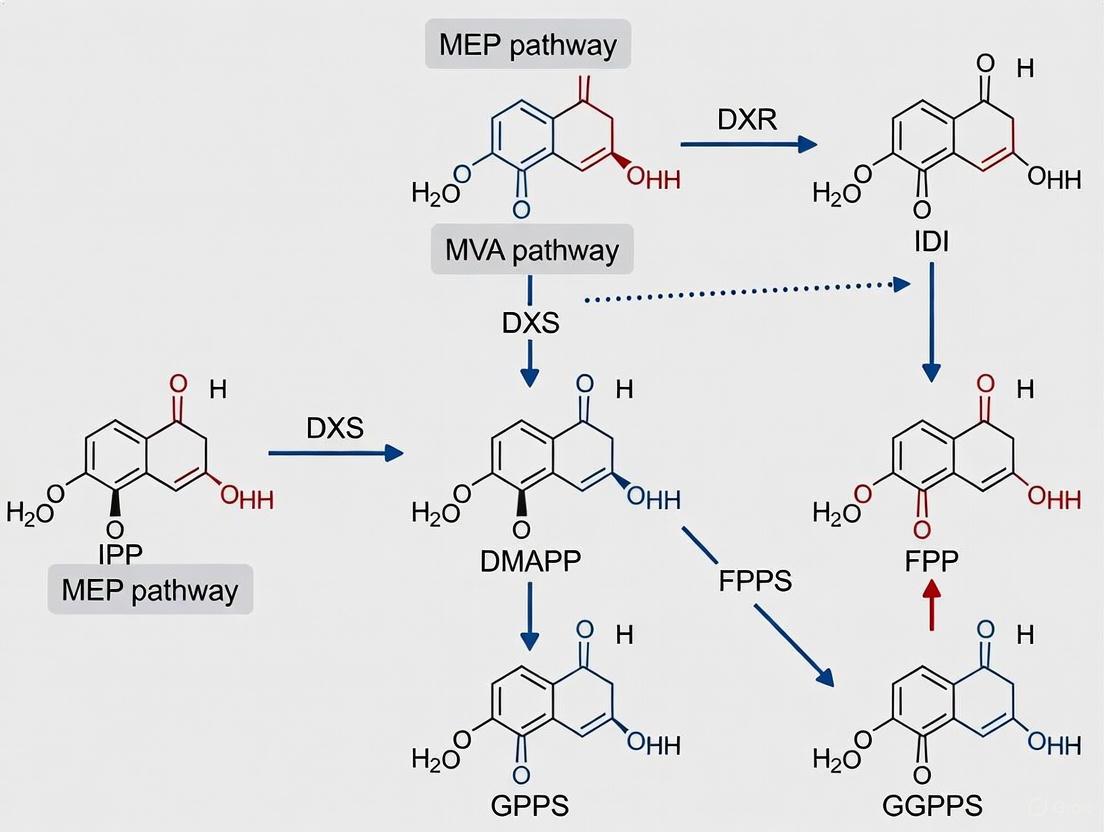
Pathway Visualization and Regulatory Networks
Figure 1: Isoprenoid Precursor Biosynthesis Pathways and Key Regulatory Nodes
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagent Solutions for Isoprenoid Precursor Studies
| Reagent/Category | Specific Examples | Function/Application | Technical Notes |
|---|---|---|---|
| Pathway Inhibitors | Fosmidomycin (DXR inhibitor) [12], Mevinolin (HMGR inhibitor) [12], Pyruvamines (TAA1/TAR inhibitors) [15] | Elucidating pathway contributions; probing enzyme essentiality | Use at optimized concentrations to avoid off-target effects; validate with rescue experiments |
| Stable Isotope Tracers | [U-¹³C]Glucose, [1-¹³C]Pyruvate, [¹⁵N]Anthranilate, [²H₃]Mevalonolactone [15] | Metabolic flux analysis; pathway utilization studies | Ensure isotopic purity; optimize labeling time for intermediate detection |
| Analytical Standards | IPP, DMAPP, Mevalonate-5-P, HMBPP, Prenyl phosphates (GPP, FPP, GGPP) [11] | LC-MS/MS quantification; method validation | Commercial availability varies; may require custom synthesis for some intermediates |
| Enzymes for IUP | Choline Kinase (ScCK), Isopentenyl Phosphate Kinase (AtIPK), IDI [12] | Constructing synthetic isoprenoid pathways | Screen orthologs for improved kinetics; consider codon optimization for heterologous expression |
| Specialized Columns | Chiral columns (e.g., CHIRALPAK IA-3), C18 reverse-phase, NH2P-50 4E [11] [16] | Separation of isomer intermediates (IPP/DMAPP); glycan analysis | Chiral columns provide baseline IPP/DMAPP separation but require careful maintenance |
| Quenching/Extraction Solutions | Cold methanol/water mixtures, Ethanol with formic acid, Chloroform:methanol:water [11] [16] | Metabolite stabilization and extraction | Optimize composition for specific microorganisms; maintain cold chain |
Concluding Remarks
The engineering of isoprenoid precursor synthesis requires a multifaceted understanding of pathway topology, regulatory constraints, and sophisticated analytical capabilities. The MEP and MVA pathways, with their complex regulation and cofactor requirements, present both challenges and opportunities for metabolic engineering. The development of synthetic pathways like the IUP demonstrates how alternative route design can bypass inherent limitations of natural systems. Emerging discoveries of non-canonical functions for pathway enzymes, such as the role of HDS in RNA editing, highlight the integrated nature of isoprenoid metabolism with broader cellular processes [14]. The experimental frameworks and reagent tools detailed herein provide a foundation for advancing research in isoprenoid pathway engineering, with significant implications for pharmaceutical development and industrial biotechnology.
Isoprenoids, also known as terpenoids, constitute the largest and most structurally diverse class of natural products with over 55,000 identified compounds, playing essential roles across all kingdoms of life [17] [18]. These compounds perform critical biological functions including membrane fluidity modulation (cholesterol, hopanoids), photoprotection (carotenoids), growth regulation (gibberellins), and defense mechanisms [6] [19]. All isoprenoids derive from two universal five-carbon building blocks: isopentenyl diphosphate (IPP) and dimethylallyl diphosphate (DMAPP) [20] [18]. Nature has evolved two distinct biosynthetic routes to produce these precursors: the mevalonate (MVA) pathway and the methylerythritol phosphate (MEP) pathway [17]. These pathways differ in their evolutionary history, subcellular localization, starting substrates, energy requirements, and distribution across organisms. Understanding the fundamental distinctions between these pathways provides the foundation for targeted metabolic engineering strategies aimed at optimizing production of high-value terpenoids for pharmaceutical, nutraceutical, and industrial applications.
Comparative Analysis of Pathway Distribution
Kingdom-Specific Pathway Utilization
The distribution of the MVA and MEP pathways across biological kingdoms reveals a fascinating evolutionary pattern, summarized in Table 1. The MVA pathway represents the ancestral route, predominantly found in Archaea, animals, and fungi, where it localizes to the cytoplasm [17] [18]. In contrast, the MEP pathway is primarily utilized by most bacteria and operates within plastids of plastid-bearing eukaryotes including plants and algae [6] [18]. Plants represent a unique case as they have retained both pathways, with the MVA pathway operating in the cytoplasm/endoplasmic reticulum and the MEP pathway functioning within plastids [20] [18]. This compartmentalization allows plants to efficiently utilize diverse carbon sources and regulate isoprenoid production in response to environmental conditions [20].
Table 1: Distribution of MVA and MEP Pathways Across Biological Kingdoms
| Kingdom/Domain | MVA Pathway | MEP Pathway | Localization | Evolutionary Origin |
|---|---|---|---|---|
| Animals | ✓ Present | ✗ Absent | Cytoplasm | Ancestral eukaryotic |
| Fungi | ✓ Present | ✗ Absent | Cytoplasm | Ancestral eukaryotic |
| Archaea | ✓ Present (often partial) | ✗ Absent | Cytoplasm | Ancestral archaeal |
| Most Bacteria | ✗ Absent (exceptions) | ✓ Present | Plastids | Bacterial endosymbionts |
| Plants | ✓ Present | ✓ Present | Cytoplasm (MVA) / Plastids (MEP) | Dual origin |
| Green Algae | ✗ Often absent | ✓ Present | Plastids | Cyanobacterial endosymbiont |
Evolutionary Origins and Phylogenetic Relationships
Genomic analyses reveal that the MEP pathway in plastid-bearing eukaryotes originated from multiple bacterial lineages through endosymbiotic gene transfer [18]. Phylogenetic evidence demonstrates that cyanobacteria contributed genes for DXR and HDR enzymes, while Chlamydiae provided CMS and CMK genes [18]. The DXS enzyme appears to have been acquired from alpha-proteobacteria [18]. This polyphyletic origin explains the complex evolutionary history and distribution patterns observed in modern organisms. Interestingly, most green algae, including Chlamydomonas reinhardtii, rely exclusively on the MEP pathway, having lost the MVA pathway during evolution [4] [21]. In contrast, diatoms maintain both pathways, suggesting divergent evolutionary trajectories among algal lineages [21].
Pathway Architecture and Regulation
MVA Pathway Biochemistry
The mevalonate pathway initiates with three molecules of acetyl-CoA, consuming 2 NADPH and 3 ATP molecules to produce one IPP molecule [4] [20]. Key enzymatic steps include the condensation of acetyl-CoA to form hydroxymethylglutaryl-CoA (HMG-CoA), followed by the rate-limiting reduction catalyzed by HMG-CoA reductase (HMGR) to produce mevalonate [4] [17]. Subsequent phosphorylation and decarboxylation steps yield IPP, which is isomerized to DMAPP by IPP isomerase (IDI) [17]. The classical MVA pathway employs phosphomevalonate kinase (PMK) and mevalonate diphosphate decarboxylase (MDD), while an alternative MVA pathway utilizes mevalonate phosphate decarboxylase (MPD) and isopentenyl phosphate kinase (IPK), which has been identified across all three domains of life [19].
MEP Pathway Biochemistry and Unique Features
The MEP pathway begins with the condensation of pyruvate and glyceraldehyde-3-phosphate (G3P), requiring 3 ATP and 3 NADPH molecules to produce both IPP and DMAPP [4] [20]. The first committed step is catalyzed by DXP synthase (DXS), considered a major metabolic bottleneck [4] [17]. A distinctive feature of the MEP pathway is the presence of two iron-sulfur (Fe-S) cluster enzymes, IspG and IspH, at the terminal steps [6]. These Fe-S clusters confer oxygen sensitivity and position the MEP pathway as an oxidative stress sensor and response system [6]. Under oxidative conditions, the intermediate MEcDP accumulates and functions as a stress-specific retrograde signaling molecule, coordinating nuclear gene expression in response to plastidial stress [6] [18].
Table 2: Comparative Analysis of MVA and MEP Pathway Characteristics
| Characteristic | MVA Pathway | MEP Pathway |
|---|---|---|
| Initial Substrates | Acetyl-CoA (3 molecules) | Pyruvate + G3P |
| Energy Cofactors Consumed | 2 NADPH + 3 ATP per IPP | 3 NADPH + 3 ATP per IPP/DMAPP |
| Theoretical Carbon Yield (Glucose) | 25.2% | 30.2% |
| Key Regulatory Enzymes | HMGR | DXS, DXR, IspG, IspH |
| Oxygen Sensitivity | Low (No Fe-S clusters in eukaryotic version) | High (Contains Fe-S cluster enzymes) |
| Pathway Intermediates with Signaling Functions | Not reported | MEcDP (stress signaling) |
Experimental Protocols for Pathway Engineering
Protocol: Engineering a Heterologous MVA Pathway in Chlamydomonas reinhardtii
Background and Principle
Green algae including Chlamydomonas reinhardtii naturally lack the MVA pathway, relying exclusively on the plastidial MEP pathway for isoprenoid precursor synthesis [4]. This protocol describes the introduction of a complete heterologous MVA pathway into C. reinhardtii to enhance isoprenoid production, resulting in a seven-fold increase in IPP levels (373.75 ± 7.176 μg/g DCW) [4].
Materials and Reagents
- Chlamydomonas reinhardtii strain UVM4
- Tris-Acetate-Phosphate (TAP) medium
- Antibiotics: zeocin (10 μg/mL), paromomycin (10 μg/mL), hygromycin (10 μg/mL)
- Optimized genes: mvaE and mvaS from Enterococcus faecalis; mvk, pmk, mvaD, and Idi from Saccharomyces cerevisiae
- Plasmid constructs for nuclear expression
- Glass beads (0.4-0.6 mm diameter) for transformation
Procedure
- Strain Cultivation: Maintain C. reinhardtii (UVM4) in TAP medium under a 16/8 h light/dark cycle at 25°C with photon fluence rate of 120 μmol m⁻² s⁻¹ [4].
- Plasmid Construction: Synthesize codon-optimized genes and clone into appropriate expression vectors containing selective markers [4].
- Two-Step Transformation:
- First Transformation: Introduce the upstream MVA module (mvaE and mvaS) using glass bead transformation method.
- Selection: Screen transformants on TAP plates supplemented with zeocin (10 μg/mL).
- Second Transformation: Introduce the downstream MVA module (mvk, pmk, mvaD, Idi) into selected upstream transformants.
- Selection: Screen double transformants on TAP plates containing both paromomycin and hygromycin (10 μg/mL each) [4].
- Analytical Verification:
- Confirm gene integration via PCR and expression via RT-qPCR.
- Quantify pathway intermediates (mevalonate, IPP) and final isoprenoid products using GC-MS or LC-MS [4].
Technical Notes
- The upstream MVA module alone can significantly enhance carbon flux toward IPP synthesis.
- Co-expression of both upstream and downstream modules is essential for complete pathway functionality.
- Optimal cultivation conditions (light, temperature, medium composition) are critical for maximizing isoprenoid yields.
Protocol: Optimizing MEP Pathway in Escherichia coli for Amorphadiene Production
Background and Principle
E. coli utilizes the native MEP pathway for isoprenoid biosynthesis. This protocol describes the enhancement of amorphadiene production through MEP pathway engineering and bioprocess optimization, achieving yields of 6.1 g/L [22].
Materials and Reagents
- E. coli production strain
- Exogenous MEP pathway genes: dxs2 from Streptomyces avermitilis, idi from Bacillus subtilis
- High-cell density fermentation system
- Controlled glucose feeding system
- Analytical standards for amorphadiene quantification
Procedure
- Strain Engineering:
- Co-express dxs2 from S. avermitilis and idi from B. subtilis in E. coli host strain.
- Verify gene expression and enzyme activity.
- Shake Flask Evaluation:
- Cultivate engineered strain in appropriate medium.
- Compare amorphadiene production with wild-type strain (expected: 12.2-fold increase) [22].
- High-Cell Density Fermentation:
- Establish fed-batch process with controlled glucose feeding.
- Recognize 24-72 h as critical period for product biosynthesis.
- Glucose Feeding Optimization:
- Optimize glucose feeding rate during 24-72 h phase (increases production from 2.5 to 4.85 g/L).
- Modulate feeding rate during exponential phase to control metabolic burden (further increases yield to 6.1 g/L) [22].
Technical Notes
- Balanced precursor supply (pyruvate and G3P) is crucial for MEP pathway efficiency.
- Combining pathway engineering with bioprocess optimization maximizes yields.
- Post-72 h metabolic attenuation requires careful feeding strategy management.
Visualization of Pathway Distribution and Engineering Workflows
Pathway Distribution Across Kingdoms
Figure 1: Distribution of MVA and MEP Pathways Across Biological Kingdoms
MVA Pathway Engineering Workflow in Chlamydomonas reinhardtii
Figure 2: MVA Pathway Engineering Workflow in C. reinhardtii
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Isoprenoid Pathway Engineering
| Reagent/Resource | Function/Application | Example Sources/Organisms |
|---|---|---|
| Codon-Optimized Genes | Heterologous expression enhancement | mvaE, mvaS (E. faecalis); mvk, pmk, mvaD, Idi (S. cerevisiae); dxs2 (S. avermitilis) [4] [22] |
| Selection Antibiotics | Transformant selection | Zeocin, paromomycin, hygromycin (10 μg/mL each) [4] |
| Analytical Standards | Metabolite quantification | Mevalonate, IPP, DMAPP, specific terpenoids (e.g., amorphadiene) [4] [22] |
| Pathway Inhibitors | Metabolic flux studies | Fosmidomycin (DXR inhibitor) [17] |
| Culture Media | Optimized microbial/algol growth | TAP medium (C. reinhardtii), M9 minimal medium (E. coli) [4] [23] |
| Expression Vectors | Gene expression in host systems | Nuclear expression vectors (C. reinhardtii), pET-based systems (E. coli) [4] [23] |
Applications in Metabolic Engineering and Biotechnology
The distinct characteristics of the MVA and MEP pathways present unique opportunities for metabolic engineering. The MEP pathway offers a higher theoretical carbon yield (30.2% on glucose) compared to the MVA pathway (25.2%), making it attractive for microbial production platforms [6]. However, engineering the MEP pathway is complicated by its oxygen sensitivity and complex regulation, particularly involving the Fe-S cluster enzymes IspG and IspH [6]. Successful engineering strategies often involve modular pathway optimization, as demonstrated by the combination of the Entner-Doudoroff pathway with the MEP pathway in E. coli, which increased isoprene production more than six-fold compared to traditional Embden-Meyerhof pathway pairing [23].
For organisms lacking one pathway, introduction of the complementary pathway has proven highly effective. In cyanobacteria, which naturally possess only the MEP pathway, introduction of a heterologous MVA pathway enhanced isoprene production 2.5-fold [4]. Similarly, engineering the MVA pathway into the green alga C. reinhardtii resulted in substantial increases in IPP levels [4]. These successes highlight the potential of pathway engineering to overcome inherent metabolic limitations and optimize isoprenoid production in diverse host organisms.
Advanced engineering approaches now focus on dynamic regulation, compartmentalization, and co-factor balancing to further enhance pathway efficiency. The discovery of the alternative MVA pathway utilizing MPD and IPK across all domains of life opens new possibilities for engineering simplified or orthogonal isoprenoid biosynthetic routes [19]. Furthermore, the recognition of the MEP pathway's role in oxidative stress signaling suggests potential engineering strategies that integrate stress response with production enhancement [6].
Isoprenoids, also known as terpenoids, represent the most complex and diverse class of natural products, with over 80,000 identified structures playing crucial roles in pharmaceutical, flavor, fragrance, and biofuel applications [24]. This structural diversity originates from a remarkably modular biosynthetic system centered on prenyl diphosphates, which serve as the universal five-carbon building blocks for constructing terpenoid skeletons [25] [24]. The sophisticated engineering of microbial cell factories now enables systematic exploitation of these pathways for both canonical terpenoid production and the creation of novel analogs with tailored properties [26]. This Application Note provides detailed methodologies and current data for engineering isoprenoid biosynthetic pathways in microbial hosts, with particular emphasis on enhancing precursor supply and controlling downstream diversification.
Biosynthetic Pathways to Prenyl Diphosphate Precursors
All terpenoid biosynthesis initiates with the formation of the fundamental C5 precursors, isopentenyl pyrophosphate (IPP) and dimethylallyl pyrophosphate (DMAPP). Microbial cell factories employ three principal pathways to generate these essential building blocks, each with distinct metabolic characteristics and engineering considerations.
Table 1: Comparison of Prenyl Diphosphate Biosynthetic Pathways
| Pathway | Key Substrates | Organismal Distribution | Carbon Efficiency | ATP Requirement | Notable Features |
|---|---|---|---|---|---|
| Methylerythritol Phosphate (MEP) | Pyruvate + G3P [10] | Most bacteria, plant plastids, cyanobacteria, microalgae [10] [27] | High (theoretically 100% from glucose) [25] | 3 ATP/IPP [5] | Oxygen-sensitive IspG/H enzymes; Complex regulation [12] |
| Mevalonate (MVA) | Acetyl-CoA [27] | Eukaryotes, archaea, certain bacteria [27] | Moderate [25] | 5 ATP/IPP (canonical) [5] | Extensive regulation; Cytotoxic intermediates [24] |
| Archaeal MVA | Acetyl-CoA [5] | Major archaeal groups [5] | Moderate | 3 ATP/IPP [5] | Energy-saving; Oxygen-sensitive PMDh enzyme [5] |
| Isopentenol Utilization (IUP) | Isoprenol/Prenol [12] [26] | Engineered systems [12] [26] | Dependent on alcohol input | 2 ATP/IPP [12] | Decoupled from central carbon metabolism; Very high flux capacity [12] |
Diagram 1: Biosynthetic Pathways to Prenyl Diphosphate Precursors. This workflow illustrates the four major routes for generating IPP and DMAPP in engineered microbial systems, highlighting key intermediates and the convergence toward the central prenyl diphosphate pool.
Engineering Enhanced Prenyl Diphosphate Supply
Optimizing the flux toward prenyl diphosphates represents the most critical step in terpenoid metabolic engineering. The following protocols and data summarize current best practices for pathway selection and enhancement.
Protocol: Implementing the Isopentenol Utilization Pathway (IUP) inE. coli
Principle: The IUP pathway bypasses native regulatory mechanisms by directly converting exogenous isopentenol isomers (isoprenol/prenol) to IPP and DMAPP via two phosphorylation steps, enabling extremely high flux decoupled from central carbon metabolism [12] [26].
Materials:
- E. coli strain KO1 (ΔispG, MEP pathway knockout) [12]
- Plasmids: pSEVA228-pro4IUPi (constitutive) or pTET-IUPi (inducible) expressing ScCK, AtIPK, and IDI [12]
- M9 minimal medium with 0.6 mM isoprenol or prenol [12]
- Anaerobic chamber for oxygen-sensitive setups
Procedure:
- Transform E. coli KO1 with IUP expression plasmids using standard heat-shock transformation.
- Plate transformed cells on M9 minimal agar containing appropriate antibiotics and 0.6 mM isoprenol.
- Incubate at 37°C for 48-72 hours until colonies appear (extended lag phase expected).
- For production cultures, inoculate 5 mL M9 medium with single colonies and grow with shaking (220 rpm) at 37°C.
- Induce IUP expression with anhydrotetracycline (100 ng/mL) for pTETIUP strains at OD600 ≈ 0.6.
- Supplement with isopentenol isomers (0.1-1% v/v) throughout cultivation period.
- Monitor growth and metabolite production over 24-96 hours.
Technical Notes:
- IUP expression can rescue MEP-knockout strains in minimal media without mevalonate supplementation [12].
- The IUP flux typically exceeds capacity of downstream pathways, requiring balancing of terpene synthase expression [12].
- Choline kinase from S. cerevisiae (ScCK) demonstrates superior activity for the first phosphorylation step compared to bacterial alternatives [12] [26].
Protocol: Employing Archaeal MVA Pathway in AerobicE. coliCultivation
Principle: The archaeal MVA pathway reduces ATP consumption by 40% compared to the canonical eukaryotic pathway, but requires special handling of oxygen-sensitive phosphomevalonate dehydratase (PMDh) [5].
Materials:
- E. coli BL21(DE3) or similar production strain
- Plasmid system expressing full archaeal MVA pathway from M. mazei (PMDh, AMPD, IPK, MVK) [5]
- LB or terrific broth with appropriate antibiotics
- Anaerobic工作站 for plasmid construction and initial testing
Procedure:
- Co-transform E. coli with plasmids encoding the complete archaeal MVA pathway and terpenoid biosynthetic enzymes (e.g., lycopene or β-farnesene pathway).
- For pre-culture, inoculate 5 mL medium and grow aerobically overnight at 30°C with shaking.
- Dilute main culture to OD600 = 0.1 in fresh medium containing 0.5% glycerol as carbon source.
- Induce pathway expression with 0.5 mM IPTG at OD600 ≈ 0.6.
- Supplement with 5 g/L mevalonolactone (hydrolyzes to MVA intracellularly) if using partial pathway.
- For enhanced PMDh stability, maintain cultures at 30°C with moderate shaking (180 rpm) to reduce oxygenation.
- Harvest cells during stationary phase for terpenoid quantification.
Technical Notes:
- Co-expression of PMDh subunits (MM1525 and MM1524 from M. mazei) is essential for functional holoenzyme [5].
- Archaeal MVA pathway function in aerobic conditions requires optimized PMDh expression and can achieve ~85% of canonical pathway productivity under optimal conditions [5].
- For highest yields, consider two-stage cultivation with initial aerobic growth followed by oxygen-limited production phase.
Table 2: Representative Terpenoid Titers Achieved via Pathway Engineering
| Target Compound | Host Organism | Engineering Strategy | Maximum Titer | Key Pathway Enzymes |
|---|---|---|---|---|
| Amorpha-4,11-diene [28] | E. coli | Balanced MK and ADS expression; MVA pathway | 7-fold increase over baseline [28] | Mevalonate kinase (MK), Amorphadiene synthase (ADS) |
| Lycopene [25] | E. coli | MAGE of 20 DXP pathway genes | 9 mg/g DCW [25] | DXP pathway enzymes, CrtEIB |
| Geraniol [25] | E. coli | Fusion tags; acyltransferase + esterase system | 13.19 g/L [25] | GGPS, Geraniol synthase, Alcohol acyltransferase |
| β-Farnesene [25] [5] | E. coli | Eukaryotic or archaeal MVA pathway | 130 g/L (in yeast) [25] | FPPS, Farnesene synthase |
| Linalool analogs [26] | S. cerevisiae | IUP with AtFKI/AtIPK + analog substrates | Varies by analog | AtFKI, AtIPK, Linalool synthase |
Diversification Strategies from Prenyl Diphosphate Intermediates
The controlled elongation and cyclization of prenyl diphosphates generates the remarkable structural diversity of terpenoids. Engineering these downstream processes requires precise manipulation of terpene synthases and prenyltransferases.
Protocol: Engineering Aromatic Prenyltransferases for Phenolic Compound Diversification
Principle: Prenyltransferases catalyze the addition of prenyl chains (C5-C20) to aromatic acceptors, dramatically altering bioactivity and membrane permeability of phenolic compounds [29].
Materials:
- Cloned prenyltransferase genes (e.g., NovQ, FgaPT2, NphB)
- Soluble aromatic substrates (flavonoids, stilbenoids, coumarins)
- Prenyl diphosphate donors (GPP, FPP, GGPP) or isopentenol for IUP integration
- E. coli or S. cerevisiae expression system with strong inducible promoters
- HPLC-MS system for product analysis
Procedure:
- Select prenyltransferase based on target phenolic substrate and desired prenyl chain length.
- Clone gene into appropriate expression vector with N-terminal solubility tags (MBP, Trx).
- Express in E. coli BL21(DE3) with 0.1 mM IPTG induction at 18°C for 16-20 hours.
- Purify enzymes using affinity chromatography corresponding to solubility tag.
- Set up 100 μL reaction mixtures containing: 50 mM Tris-HCl (pH 7.5), 5 mM MgCl₂, 0.1 mM prenyl diphosphate, 0.2 mM aromatic acceptor, and purified enzyme.
- Incubate at 30°C for 2-4 hours with gentle shaking.
- Extract products with ethyl acetate and analyze by HPLC-MS.
- For whole-cell biotransformation, supplement culture medium with 0.1-1 mM aromatic substrates during induction phase.
Technical Notes:
- Membrane-associated plant PTs often require yeast expression systems for proper folding [29].
- Directed evolution of PTs using structure-guided approaches can enhance catalytic efficiency and substrate specificity [29].
- Combinatorial biosynthesis with IUP enables production of novel prenylated analogs using non-canonical prenyl donors [26].
Diagram 2: Diversification Pathways from Central Prenyl Diphosphate Intermediates. This workflow illustrates the enzymatic transformation of basic prenyl diphosphate building blocks into structurally diverse terpenoid classes through controlled chain elongation, cyclization, and functionalization reactions.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for Isoprenoid Pathway Engineering
| Reagent Category | Specific Examples | Function & Application | Key Characteristics |
|---|---|---|---|
| Kinases for IUP | ScCK (S. cerevisiae), AtFKI (A. thaliana) [12] [26] | First phosphorylation step (alcohol → monophosphate) | ScCK has broad substrate specificity; AtFKI shows higher efficiency in yeast [26] |
| Second-step Kinases | AtIPK (A. thaliana) [12] [26] | Second phosphorylation (monophosphate → diphosphate) | High kcat/KM; Compatible with non-canonical substrates [12] |
| Prenyltransferases | NovQ, FgaPT2, NphB [29] | Transfer prenyl groups to aromatic acceptors | Varying chain length specificity (GPP, FPP, GGPP); Often membrane-associated [29] |
| Terpene Synthases | Amorphadiene synthase, Linalool synthase, Farnesene synthase [25] [28] | Cyclize/couple prenyl diphosphates to form terpene skeletons | Often produce multiple products; Engineering required for specificity [24] |
| Pathway Enzymes | DXS, IDI, FPPS, GGPPS [25] [24] | Native MEP/MVA pathway and chain elongation | DXS often rate-limiting in MEP pathway; IDI balances IPP/DMAPP ratio [24] |
| Host Strains | E. coli KO1 (ΔispG), EGY48, BL21(DE3) variants [12] [5] | Production chassis with optimized metabolism | Specialized for pathway rescue (KO1) or high protein expression (BL21) [12] [5] |
The systematic engineering of prenyl diphosphate biosynthesis represents the foundational step in reprogramming microbial factories for terpenoid production. The development of orthogonal systems like the IUP pathway and energy-efficient archaeal MVA variants provides increasingly sophisticated tools for overcoming native regulatory constraints [12] [5]. Future directions will focus on dynamic pathway regulation to balance precursor supply with downstream capacity, machine learning-assisted enzyme engineering for precise product specificity, and integration of non-canonical building blocks to access novel chemical space [26] [29]. The protocols and data presented herein provide a robust foundation for advancing these efforts toward the sustainable production of both natural terpenoids and designed analogs with tailored properties.
Engineering Blueprints: Strategies for Optimizing Isoprenoid Production
Within the field of isoprenoid biosynthetic pathway engineering, the selection of an appropriate microbial host is a foundational decision that profoundly influences the success and scalability of production. Isoprenoids, also known as terpenoids, constitute the largest class of natural products with over 70,000 identified compounds, serving critical roles in pharmaceuticals, nutraceuticals, fragrances, and advanced biofuels [30]. For decades, metabolic engineering efforts have predominantly relied on conventional model organisms, namely Escherichia coli and Saccharomyces cerevisiae, leveraging their well-characterized genetics and rapid growth kinetics [31]. However, the commercial viability of microbial isoprenoid production is often constrained by precursor limitations, high substrate costs, and the inherent metabolic burdens placed on these hosts [30]. This application note delineates the strategic evolution in host selection, comparing the established paradigms of conventional hosts with the emerging potential of non-model organisms, and provides detailed experimental protocols to guide research in this rapidly advancing field. The overarching goal is to equip researchers with the practical knowledge to evaluate and implement the most suitable host platform for their specific isoprenoid production targets, framed within the context of a comprehensive thesis on pathway engineering.
Conventional Microbial Hosts: Established Workhorses
1Escherichia coliandSaccharomyces cerevisiae
The well-defined physiology and extensive genetic toolkits available for E. coli and S. cerevisiae have cemented their status as the primary hosts for heterologous isoprenoid production. Table 1 summarizes the core characteristics of these two conventional workhorses. E. coli utilizes the native methylerythritol phosphate (MEP) pathway for the synthesis of the universal C5 isoprenoid precursors, isopentenyl diphosphate (IPP) and dimethylallyl diphosphate (DMAPP). This pathway offers a higher theoretical carbon yield (0.83 C-mol/C-mol) from glucose compared to the mevalonate (MVA) pathway [31]. In contrast, S. cerevisiae operates the native MVA pathway, which is more energy-demanding but provides a cytosolic environment conducive to the functional expression of eukaryotic enzymes, particularly cytochrome P450s (CYPs) that are often required for the decoration of complex terpenoid skeletons [32] [31].
Table 1: Characteristics of Conventional Isoprenoid Production Hosts
| Feature | Escherichia coli | Saccharomyces cerevisiae |
|---|---|---|
| Native Pathway | MEP Pathway | MVA Pathway |
| Theoretical Carbon Yield (from Glucose) | 0.83 C-mol/C-mol [31] | 0.56 C-mol/C-mol [31] |
| Key Precursors | Pyruvate & Glyceraldehyde-3-phosphate [10] | Acetyl-CoA [10] |
| Genetic Tools | Extensive, well-developed | Extensive, well-developed |
| Key Advantage | High theoretical yield; rapid growth | Superior for expressing eukaryotic P450s; compartmentalization |
| Primary Challenge | Lack of native P450 redox systems; precursor toxicity | Competing pathways (e.g., ergosterol synthesis); lower theoretical yield |
Experimental Protocol: Engineering a Base Isoprenoid Platform inE. coli
The following protocol details the creation of a high-yielding sesquiterpene platform in E. coli, a common strategy for proof-of-concept and initial production studies.
- Objective: To engineer an E. coli strain for the overproduction of a sesquiterpene (e.g., bisabolene) from glucose.
- Principle: The strategy involves two key steps: (1) augmenting the flux of the native MEP pathway toward IPP and DMAPP, and (2) introducing a heterologous terpene synthase to convert the intermediate farnesyl pyrophosphate (FPP) into the target molecule.
Materials:
- Strains: E. coli MG1655 or BL21(DE3)
- Vectors: A system for chromosomal integration or plasmid-based expression (e.g., pET series with T7 promoter)
- Key Genes for Pathway Engineering:
- dxs (1-deoxy-D-xylulose-5-phosphate synthase)
- idi (IPP isomerase)
- ispDF (operon for MEP pathway enzymes)
- Heterologous ERG20 (FPP synthase) from yeast, codon-optimized
- Terpene synthase gene (e.g., bisB for bisabolene), codon-optimized
Procedure:
- Strain Construction: a. Modulate the MEP pathway: Clone the dxs and idi genes under the control of a strong, inducible promoter (e.g., PT7) on a plasmid. Alternatively, integrate strong, constitutive promoters upstream of the genomic dxs and idi loci. b. Amplify FPP pool: Introduce a plasmid expressing a heterologous ERG20 (FPP synthase) to enhance the condensation of IPP and DMAPP into FPP. c. Introduce product synthase: Co-transform the strain with a second plasmid expressing the codon-optimized terpene synthase gene (e.g., bisB) under an inducible promoter.
- Fermentation and Analysis: a. Inoculate 5 mL of LB medium with antibiotics and grow overnight at 37°C. b. Sub-culture into 50 mL of defined mineral medium (e.g., M9) with 2% glucose and appropriate antibiotics in a 250 mL baffled flask. c. Incubate at 30-37°C with shaking at 250 rpm. Induce gene expression with IPTG (e.g., 0.1-0.5 mM) when the culture reaches mid-exponential phase (OD600 ≈ 0.6). d. Continue cultivation for 24-72 hours post-induction. For hydrophobic terpenes, consider adding a solvent overlay (e.g., 10% dodecane) to capture volatilized products and mitigate toxicity. e. Analyze production via GC-MS or HPLC. Extract cells and supernatant (or the solvent overlay) with ethyl acetate, concentrate the organic phase under nitrogen gas, and resuspend in an appropriate solvent for instrumental analysis.
The Rationale for Non-Model Organisms
While E. coli and S. cerevisiae have achieved remarkable production titers for some isoprenoids, their limitations have spurred the exploration of non-model hosts. These organisms often possess innate physiological attributes that can circumvent key bottlenecks. Microalgae, for instance, are photosynthetic organisms that can fix atmospheric CO₂, eliminating the need for expensive organic carbon sources [10]. Many microalgae also possess native MVA or MEP pathways and naturally produce a diverse array of isoprenoids (e.g., carotenoids), reducing the need for extensive pathway engineering [10]. Oleaginous yeasts like Yarrowia lipolytica and Rhodosporidium toruloides are characterized by an abundant acetyl-CoA pool, the primary precursor for the MVA pathway, and a natural tolerance to high levels of lipophilic compounds, which is beneficial for terpenoid production [31] [30]. Furthermore, certain non-model bacteria and archaea exhibit extreme stress tolerance and, crucially, the ability to utilize unconventional, low-cost carbon substrates such as methane, methanol, and synthesis gas (syngas) [30]. This capacity to valorize waste carbon streams presents a compelling strategy for dramatically reducing production costs, which are currently dominated by feedstock expenses [30].
Table 2: Promising Non-Model Organisms for Isoprenoid Production
| Organism | Native Pathway | Key Innate Advantage | Potential Application |
|---|---|---|---|
| Microalgae (e.g., Phaeodactylum tricornutum) | MEP and/or MVA [10] | CO₂ fixation; photoautotrophic growth; natural high-value isoprenoid producer [10] | Sustainable production of carotenoids and biofuels |
| Oleaginous Yeast (e.g., Yarrowia lipolytica) | MVA [31] | Abundant acetyl-CoA pool; high lipid and terpenoid storage capacity [31] [30] | High-titer production of sesquiterpenes and triterpenes |
| Rhodobacter spp. (Purple non-sulfur bacteria) | MEP [30] | Metabolic versatility: phototrophic, mixotrophic, or chemotrophic growth on diverse carbon sources [30] | Conversion of agro-industrial wastes to carotenoids |
| Methylotrophic Bacteria (e.g., Methylobacterium extorquens) | MEP or MVA [30] | Utilizes methanol and other C1 compounds as carbon source [30] | Low-cost production platform using methane/methanol |
Advanced Strategies: Synthetic Microbial Consortia
A sophisticated approach to overcome the limitations of individual hosts is the use of synthetic microbial consortia, where pathway modules are distributed between two or more organisms [32]. A seminal example is the co-culture of E. coli and S. cerevisiae for the production of oxygenated taxol precursors [32]. In this system, E. coli was engineered to produce taxadiene efficiently, while S. cerevisiae, a superior host for functional P450 expression, performed the oxygenation steps. To ensure stable co-culture, a mutualistic relationship was engineered: the carbon source was switched to xylose, which E. coli metabolizes excretes acetate, and S. cerevisiae in turn consumes the acetate without producing inhibitory ethanol [32]. This strategy synergistically combines the unique strengths of both conventional hosts.
Experimental Protocol: Establishing a Mutualistic Co-culture for Pathway Division
- Objective: To produce an oxygenated terpenoid (e.g., taxadiene-5α-ol) using a stable co-culture of engineered E. coli and S. cerevisiae.
- Principle: The biosynthetic pathway is split. The first module (e.g., taxadiene production) is housed in E. coli, and the second module (e.g., P450-mediated oxygenation) is housed in yeast. A mutualistic relationship based on carbon and nitrogen exchange ensures population stability.
Materials:
- Strain A: E. coli engineered with the MVA pathway and taxadiene synthase.
- Strain B: S. cerevisiae engineered to express taxadiene-5α-hydroxylase and its cognate reductase.
- Medium: Defined minimal medium with xylose as the sole carbon source.
Procedure:
- Strain Preparation: Engineer the E. coli strain for high-level taxadiene production on xylose. Engineer the yeast strain with a strong promoter (e.g., UAS-GPDp) driving the P450 system [32].
- Inoculum and Reactor Setup: a. Grow pure cultures of both engineered strains overnight. b. Inoculate a bioreactor containing minimal medium with xylose. The initial inoculum ratio should be optimized (e.g., a higher initial yeast ratio may be needed to prevent acetate accumulation) [32]. c. Maintain the co-culture in fed-batch mode, with periodic feeding of xylose and essential nutrients (nitrogen, phosphorous) to support sustained growth of both populations.
- Monitoring and Analysis: a. Monitor cell density of both species by plating on selective media or using species-specific qPCR. b. Track metabolite levels (xylose, acetate, ethanol) via HPLC. c. Quantify intermediate (taxadiene) and final oxygenated product via GC-MS or LC-MS, extracting from both the culture broth and cell pellets.
The following workflow diagram illustrates the logical structure of this co-culture system.
Diagram 1: Workflow for establishing a mutualistic microbial consortium for distributed isoprenoid biosynthesis.
The Scientist's Toolkit: Essential Reagents and Solutions
Table 3: Key Research Reagent Solutions for Isoprenoid Pathway Engineering
| Reagent / Tool Category | Specific Examples | Function / Application |
|---|---|---|
| Vector Systems | pET vectors (for E. coli), pESC vectors (for S. cerevisiae), integrative plasmids | Heterologous gene expression; stable genomic integration |
| Promoters | T7, Ptrc (for E. coli); TEF1, GPD, UAS-GPD (for S. cerevisiae) [32] | Constitutive or inducible control of gene expression strength |
| Pathway Genes | dxs, idi, ispDF (MEP); ERG10, ERG13, tHMG1 (MVA); ERG20 (FPP synthase) | Augmenting precursor (IPP/DMAPP) supply |
| Analytical Standards | IPP, DMAPP, GPP, FPP, GGPP; target terpenoids (e.g., taxadiene, bisabolene) | Quantification of intermediates and products via GC-MS/LC-MS |
| Culture Additives | Dodecane (overlay), mevalonolactone (MVA precursor feed) | In-situ product extraction; precursor supplementation |
| Gene Editing Tools | CRISPR-Cas9 systems (for both model and non-model hosts) [33] | Precise genome editing, gene knock-outs, and regulatory tuning |
Visualization of Metabolic Context and Engineering Strategies
The core metabolic pathways for isoprenoid biosynthesis and key engineering strategies are visualized below. The MEP and MVA pathways converge on the universal precursors IPP and DMAPP, which are then assembled into a diverse array of terpenoid skeletons.
Diagram 2: The core metabolic map of isoprenoid biosynthesis, highlighting the MEP and MVA pathways and downstream diversification.
The strategic landscape of host selection for isoprenoid pathway engineering is expanding beyond the traditional confines of E. coli and S. cerevisiae. While these conventional hosts remain indispensable for their genetic tractability and serve as powerful platforms for pathway prototyping and validation, their inherent metabolic and physiological constraints are driving the field toward more specialized organisms. The future of sustainable and economically viable isoprenoid production lies in matching the unique chemical profile of the target molecule with the innate advantages of a particular host—be it the photosynthetic capability of microalgae, the acetyl-CoA abundance in oleaginous yeast, or the capacity of methylotrophs to consume C1 waste streams. Furthermore, advanced strategies like synthetic consortia demonstrate that the division of labor between different microbial specialists can unlock complex biosynthetic routes that are challenging for a single organism. As genetic tools for non-model hosts continue to mature, the paradigm will shift from engineering pathways into generic hosts to strategically selecting and optimizing the native capabilities of the most physiologically suited organism for the task.
In the broader context of isoprenoid biosynthetic pathway engineering, overcoming the limitation of precursor supply is a cornerstone challenge for achieving high-yield production of valuable compounds. Isoprenoids, the largest class of natural products with over 65,000 members, all originate from the universal C5 precursors isopentenyl diphosphate (IPP) and dimethylallyl diphosphate (DMAPP) [10] [34]. The inherent regulation of native metabolic pathways and the presence of rate-limiting enzymatic steps tightly constrain the flux toward these foundational building blocks, creating a fundamental bottleneck across production platforms [34]. Precursor pool enhancement encompasses a suite of metabolic engineering strategies designed to systematically identify and overcome these bottlenecks, thereby increasing the carbon flux through the pathway and enabling higher titers of target isoprenoids. This approach is universally applicable, whether engineering microbial chassis like E. coli and S. cerevisiae, optimizing native medicinal plants, or utilizing heterologous plant hosts such as Nicotiana benthamiana [35].
Core Strategies for Precursor Enhancement
Five principal strategies have emerged as the foundation for overcoming precursor limitations in isoprenoid biosynthesis [10] [34]. These approaches can be employed individually or in combination to rewire microbial, plant, or cell-free metabolism for enhanced production.
Table 1: Core Strategies for Overcoming Precursor Limitations
| Strategy | Core Principle | Key Enzymatic Targets | Exemplary Application |
|---|---|---|---|
| Overcoming Rate-Limiting Enzymes | Overexpress enzymes that catalyze the slowest, most regulated steps in the pathway to remove flux constraints [34]. | DXS (MEP pathway); HMGR, MK (MVA pathway) [34] [35]. | Overexpression of HMGR in Artemisia annua increased artemisinin yield by 22.5-38.9% [35]. |
| Expressing Heterologous Pathways | Introduce complete, often less-regulated, heterologous pathways to bypass native route or supplement precursor supply [10] [12]. | The entire MVA pathway in MEP-native hosts, or the synthetic Isopentenol Utilization Pathway (IUP) [12]. | The two-step IUP using ScCK and AtIPK sustained very high isoprenoid flux, decoupled from central carbon metabolism [12]. |
| Enhancing Cofactor Supply | Ensure adequate supply of essential cofactors (ATP, NADPH) that are consumed in large quantities during precursor synthesis [10] [36]. | Enzymes for ATP regeneration; enzymes for NADPH synthesis [36]. | In vitro lower MVA pathway required a 1:4 ratio of mevalonate to ATP for optimal isoprene production, exceeding the theoretical 1:3 consumption [36]. |
| Downregulating Competing Pathways | Knock out or knock down genes in pathways that consume IPP, DMAPP, or key intermediates, redirecting flux to the desired product [10] [34]. | IDI; prenyltransferases for non-target isoprenoids [34]. | CRISPR-Cas9-mediated knockout of competing pathways in plants enhances target terpenoid production [35]. |
| Subcellular Compartmentalization | Target pathway enzymes to specific cellular organelles (e.g., chloroplasts) to leverage localized high precursor concentrations and avoid metabolic cross-talk [10] [37]. | Chloroplast-targeting peptides for enzymes like Taxadiene Synthase (TS) and Taxadiene-5α-Hydroxylase (T5αH) [37]. | Chloroplastic compartmentalization in N. benthamiana enabled production of 56.6 μg g⁻¹ FW taxadiene and 1.3 μg g⁻¹ FW taxadiene-5α-ol [37]. |
Detailed Experimental Protocols
Protocol 1: Enhancing Flux via Rate-Limiting Enzyme Overexpression in a Microbial Host
This protocol details the process of increasing precursor flux in E. coli by overexpressing the rate-limiting enzyme 1-deoxy-D-xylulose-5-phosphate synthase (DXS) from the methylerythritol phosphate (MEP) pathway. The general workflow is as follows.
Research Reagent Solutions
Table 2: Key Reagents for Microbial Pathway Engineering
| Reagent / Material | Function / Application | Exemplary Details |
|---|---|---|
| DXS Gene Sequence | Catalyzes the first committed, rate-limiting step of the MEP pathway: condensation of pyruvate and G3P to form DXP [34]. | Codon-optimized for E. coli expression; cloned under a strong, inducible promoter (e.g., T7, pTrc). |
| Expression Plasmid | Vector for harboring and expressing the target gene in the microbial host. | A high-copy-number plasmid (e.g., pET series) with a selectable marker (e.g., kanamycin resistance). |
| E. coli Production Chassis | Engineered host strain for isoprenoid production. | Often a strain with an enhanced supply of G3P/pyruvate and/or a deleted competing pathway [34]. |
| Lycopene Reporter System | A visual and quantifiable phenotypic marker for flux through the isoprenoid pathway [12]. | Plasmid(s) expressing CrtE, CrtI, and CrtB genes, which convert FPP to lycopene. |
| Analytical Standards | For accurate quantification of metabolites and products. | IPP, DMAPP, mevalonate, and target isoprenoid (e.g., lycopene) for HPLC or GC-MS calibration. |
Step-by-Step Procedure
Gene Cloning and Strain Construction
- Amplify the coding sequence of the dxs gene using PCR primers designed for your chosen expression vector.
- Ligate the purified dxs PCR product into the plasmid using Gibson Assembly or a similar method.
- Transform the constructed plasmid into a competent E. coli production strain. If using a lycopene reporter, ensure the strain already contains the necessary crt genes on a separate, compatible plasmid or integrated into the genome.
- Plate the transformation on LB agar containing the appropriate antibiotic(s) and incubate overnight at 37°C.
Small-Scale Screening and Cultivation
- Inoculate 3-5 mL of LB medium containing antibiotics with single colonies of both the engineered strain (with dxs) and an empty vector control strain.
- Grow the cultures at 37°C with shaking until the OD600 reaches approximately 0.6.
- Induce gene expression by adding a defined concentration of inducer (e.g., 0.1-1.0 mM IPTG for pTrc or T7 promoters).
- Shift the temperature to 30°C or lower if necessary to improve protein solubility and continue incubation for 16-24 hours.
Metabolic Analysis and Flux Assessment
- Visual Screening: If a lycopene reporter is used, visually compare the color intensity of the cell pellets from the engineered and control strains after centrifugation. A deeper red color indicates higher flux through the isoprenoid pathway [12].
- Product Quantification:
- For lycopene: Harvest cells by centrifugation. Extract lycopene from the cell pellet using acetone or a methanol:acetone mixture (1:1, v/v). Measure the absorbance of the supernatant at 472 nm and calculate the concentration using a standard curve [12].
- For other isoprenoids: Analyze culture extracts using High-Performance Liquid Chromatography (HPLC) or Gas Chromatography-Mass Spectrometry (GC-MS) against authentic standards.
- Precursor Analysis: Quantify intracellular levels of pathway intermediates (e.g., DXP, MEP, IPP, DMAPP) using LC-MS/MS. This requires a rapid quenching of metabolism (e.g., with cold methanol) and specialized extraction protocols.
Protocol 2: Compartmentalized Pathway Engineering in a Plant Host
This protocol describes the enhancement of diterpenoid precursor pools in Nicotiana benthamiana by transiently expressing pathway enzymes targeted to the chloroplast, as demonstrated for taxadiene production [37].
Research Reagent Solutions
Table 3: Key Reagents for Plant Chloroplastic Engineering
| Reagent / Material | Function / Application | Exemplary Details |
|---|---|---|
| Agrobacterium tumefaciens GV3101 | Vehicle for transient gene expression in plants via agroinfiltration. | Strain transformed with the expression vectors of interest. |
| Chloroplast-Targeted Expression Vectors | Plasmids for high-level transient expression of enzymes fused to a chloroplast transit peptide. | Vectors like pEAQ-HT, which uses a CPMV-HT system for high yield. Genes (e.g., TS, T5αH) are fused to the N-terminal chloroplast transit peptide of the Rubisco small subunit [37]. |
| 4-5 Week Old N. benthamiana Plants | High-biomass, model plant host for transient expression. | Grown under standard greenhouse conditions (e.g., 16h light/8h dark, 25°C). |
| Induction Buffer | Solution for preparing Agrobacterium cultures for infiltration. | Typically contains 10 mM MES, 10 mM MgCl₂, and 150 µM acetosyringone, pH 5.6. |
| GC-MS / HPLC System | For identification and quantification of terpenoid products. | GC-MS is ideal for volatile compounds like taxadiene; HPLC for less volatile or more polar derivatives. |
Step-by-Step Procedure
Vector Construction and Agrobacterium Preparation
- Clone the genes of interest (e.g., Taxadiene Synthase - TS, Taxadiene-5α-Hydroxylase - T5αH, Cytochrome P450 Reductase - CPR) into plant expression vectors, ensuring they are in-frame with an N-terminal chloroplast transit peptide sequence.
- Transform the constructed vectors into Agrobacterium tumefaciens GV3101.
- For co-expression, inoculate single colonies of Agrobacterium harboring each plasmid into LB medium with appropriate antibiotics and 50 µM acetosyringone. Grow overnight at 28°C with shaking.
- Pellet the cultures and resuspend them in induction buffer to a final OD600 of ~0.5-1.0 for each strain. Incubate at room temperature for 2-4 hours.
- Mix the bacterial suspensions in the desired ratio (e.g., 1:1:1 for TS, T5αH, and CPR).
Plant Infiltration and Incubation
- Using a needleless syringe, gently infiltrate the mixed Agrobacterium suspension into the abaxial side of fully expanded leaves of 4-5 week old N. benthamiana plants.
- Mark the infiltrated areas. Return the plants to the greenhouse and maintain them for 4-7 days post-infiltration (dpi).
Metabolite Extraction and Analysis
- At 5 dpi, harvest leaf discs from the infiltrated zones. Fresh weight should be recorded immediately.
- For taxadiene extraction, homogenize the tissue in n-hexane or a similar non-polar solvent. Internal standards (e.g., caryophyllene) can be added for quantification.
- Analyze the organic extract by GC-MS.
- GC Conditions: Use a non-polar column (e.g., DB-5). Oven program: 50°C for 2 min, ramp to 280°C at 15°C/min, hold for 5 min.
- MS Detection: Use electron impact (EI) ionization at 70 eV. Identify taxadiene by its characteristic retention time (~18.8 min) and mass fragmentation pattern (major ions at m/z 272, 257, 122) [37].
- Quantify the product using a standard curve generated from authentic taxadiene or a structurally similar compound.
Protocol 3: Optimizing a Synthetic Pathway in a Cell-Free System
This protocol outlines the quantitative balancing of enzyme levels in a cell-free system for the high-yield conversion of mevalonate to isoprene, demonstrating precise control over precursor utilization [36].
Research Reagent Solutions
Table 4: Key Reagents for Cell-Free Pathway Optimization
| Reagent / Material | Function / Application | Exemplary Details |
|---|---|---|
| Purified Lower MVA Enzymes | Catalyze the conversion of mevalonate to isoprene. | Mevalonate kinase (MVK), phosphomevalonate kinase (PMK), diphosphomevalonate decarboxylase (MVD), isopentenyl diphosphate isomerase (IDI), isoprene synthase (ISPS). Purified from recombinant E. coli [36]. |
| Substrate & Cofactors | Raw materials and energy for the enzymatic reaction. | Sodium mevalonate; ATP; MgCl₂ (as a Mg²⁺ source, essential for kinase activity). |
| In Vitro Reaction Buffer | Provides optimal pH and ionic conditions for enzyme activity. | e.g., 50 mM Tris-HCl buffer (pH 7.5), 10 mM MgCl₂, 5% (w/v) glycerol. |
| Sealed Vial / Bioreactor | To contain the volatile isoprene product and prevent its loss. | Glass vials with PTFE/silicone septa; allows for headspace sampling. |
| GC-FID System | For sensitive and accurate quantification of gaseous isoprene. | Equipped with a GS-GasPro column or similar. |
Step-by-Step Procedure
Enzyme Expression and Purification
- Express His-tagged versions of MVK, PMK, MVD, IDI, and ISPS in E. coli. Induce expression with IPTG at a low temperature (e.g., 20°C) to improve soluble protein yield.
- Purify each enzyme using immobilized metal affinity chromatography (IMAC). Determine protein concentration and confirm purity via SDS-PAGE.
Bottleneck Identification and Unit Balancing
- Set up a series of 2 mL in vitro reactions containing 2.5 mM mevalonate, 12 mM ATP, and reaction buffer.
- Keep the concentration of four enzymes constant at 0.5 µM, while varying the concentration of the fifth enzyme from 0.02 to 5 µM. Incubate at 30-37°C.
- After a set time (e.g., 4 hours), measure isoprene production via GC. This identifies which enzymes are flux-limiting (e.g., IDI and ISPS were found to be critical bottlenecks) [36].
Optimized Reaction Assembly and Scaling
- Based on the results, establish a balanced enzyme unit. The study found an optimal molar ratio of MVK:PMK:MVD:IDI:ISPS = 1:1:1:2:16 [36].
- For a 2 mL reaction, use 5.0 µM MVK, PMK, MVD; 10.0 µM IDI; and 80.0 µM ISPS with 2.5 mM mevalonate and 10 mM ATP.
- For scale-up, proportionally reduce enzyme concentrations. A 50 mL reaction using 0.5 µM MVK, PMK, MVD; 1.0 µM IDI; and 8.0 µM ISPS can be run for extended periods (e.g., 40 hours) with continuous or batch-fed substrate addition.
Product Quantification
- Quantify isoprene production by sampling the headspace of the sealed reaction vials using a gas-tight syringe and injecting into a GC equipped with a Flame Ionization Detector (FID) or a Mass Spectrometer (MS).
- Calculate the production rate (e.g., µmol/L/h or mg/L/h) and total titer. The optimized system achieved a production rate of 6323.5 µmol/L/h (430 mg/L/h) in a 2 mL system [36].
Expanding the Carbon Menu: Engineering C1 and Waste Stream Utilization
The sustainable production of high-value isoprenoids necessitates a strategic shift from conventional sugar-based feedstocks to underutilized C1 carbon sources and industrial waste streams. This application note provides a detailed experimental framework for implementing the Isopentenol Utilization Pathway (IUP), a synthetic two-step pathway that decouples isoprenoid biosynthesis from central carbon metabolism [12] [38]. We present validated protocols for pathway construction in E. coli, quantitative analysis of metabolite flux, and strategies for host engineering to enhance precursor supply from non-conventional substrates. Designed for researchers and scientists in metabolic engineering and drug development, these protocols enable robust and high-flux production of isoprenoid precursors, aligning biomanufacturing with circular bioeconomy principles.
Isoprenoids represent one of the largest classes of natural compounds, with over 50,000 structures possessing diverse applications in pharmaceuticals, nutraceuticals, flavors, and fragrances [12]. A significant challenge in their commercial production lies in the inherent limitations of native biosynthetic pathways—the mevalonate (MVA) and methylerythritol phosphate (MEP) pathways. These pathways are inextricably linked to central carbon metabolism, suffer from complex regulatory feedback mechanisms, and compete with other cellular processes for resources [12] [39].
Expanding the "carbon menu" to include C1 compounds (e.g., methanol, formate) and waste streams (e.g., agro-industrial residues, syngas) requires metabolic architectures that are both flexible and efficient. The IUP serves as an ideal platform for this expansion. By using externally provided isopentenol isomers as substrates, which can be derived from C1-assimilating organisms or from waste carbon, the IUP bypasses the lengthy, regulated native pathways, requiring only two enzymatic steps and ATP to generate the universal isoprenoid precursors, isopentenyl diphosphate (IPP) and dimethylallyl diphosphate (DMAPP) [12]. This document details the application of this pathway for sustainable isoprenoid biosynthesis.
Background and Scientific Context
Native Isoprenoid Pathways and Their Limitations
In conventional microbial production, IPP and DMAPP are synthesized via one of two natural routes:
- The Mevalonate (MVA) Pathway: A seven-step pathway starting from acetyl-CoA, primarily found in eukaryotes and archaea [12] [13].
- The Methylerythritol Phosphate (MEP) Pathway: A seven-step pathway starting from pyruvate and glyceraldehyde-3-phosphate (G3P), found in most bacteria and the plastids of plants and microalgae [10] [12] [39].
Both pathways present engineering challenges, including susceptibility to feedback inhibition by downstream intermediates (e.g., IPP inhibition of DXS in the MEP pathway), imbalances in precursor supply, and oxygen sensitivity of iron-sulfur enzymes in the MEP pathway [12]. Furthermore, their direct connection to core metabolism complicates redirecting flux from novel carbon sources like C1 compounds.
The Engineered Isopentenol Utilization Pathway (IUP)
The IUP is a synthetic pathway that radically simplifies precursor biosynthesis (Fig. 1). It utilizes isoprenol or prenol as a substrate, which are phosphorylated in two sequential steps to form IPP and DMAPP, respectively [12]. This design offers distinct advantages for utilizing alternative carbon streams:
- Decoupling from Central Metabolism: IUP flux is independent of glycolytic or C1-assimilation intermediates, removing a major metabolic bottleneck [12].
- High Flux Capacity: The IUP has demonstrated competitive flux compared to highly optimized native pathways, sustaining high yields of downstream isoprenoids like lycopene [12] [38].
- Simplified Cofactor Requirement: The pathway requires only ATP, avoiding the complex NADPH/NADH and CTP demands of native routes [12].
Figure 1. The Two-Step Isopentenol Utilization Pathway (IUP). Isoprenol and prenol are converted to the central isoprenoid precursors IPP and DMAPP via sequential phosphorylation by choline kinase (ScCK) and isopentenyl phosphate kinase (AtIPK). Isopentenyl diphosphate isomerase (IDI) balances the IPP/DMAPP ratio.
Application Notes: Quantitative Performance of Isoprenoid Pathways
The performance of the IUP has been quantitatively compared to engineered native pathways. The table below summarizes key metrics for isoprenoid production, highlighting the efficiency of the IUP.
Table 1: Comparative Performance of Engineered Isoprenoid Biosynthesis Pathways
| Pathway / Host System | Target Product | Key Engineering Strategy | Reported Yield / Titer | Key Advantage |
|---|---|---|---|---|
| IUP in E. coli [12] | Lycopene (C40) | Expression of ScCK + AtIPK with 0.6 mM isoprenol feed | Flux competitive with highest reported native pathways | Decoupled from central carbon metabolism; simplified regulation |
| MEP in P. tricornutum [10] | Fucoxanthin | Overexpression of rate-limiting DXS enzyme | 2.4-fold increase vs. wild-type | Native pathway in industrially relevant microalgae; leverages CO2 |
| MEP/MVA Crosstalk in Diatoms [39] | Sterols, HBIs | Native complementarity of cytosolic (MVA) and plastid (MEP) pathways | Species-specific precursor allocation | Metabolic flexibility; potential for engineering novel carbon routing from C1 |
Experimental Protocols
Protocol 1: Constructing and Testing the IUP in anE. coliKnockout Host
This protocol outlines the creation of an MEP-pathway knockout strain and its rescue via the IUP, providing a direct functional assay for pathway activity [12].
Research Reagent Solutions
| Reagent / Material | Function / Description | Source / Example |
|---|---|---|
| Choline Kinase (ScCK) | Catalyzes the first phosphorylation of isoprenol/prenol to IP/DMAP. | Saccharomyces cerevisiae [12] |
| Isopentenyl Phosphate Kinase (AtIPK) | Catalyzes the second phosphorylation of IP/DMAP to IPP/DMAPP. | Arabidopsis thaliana [12] |
| Isopentenyl Diphosphate Isomerase (IDI) | Balances the cellular ratio of IPP and DMAPP. | E. coli or heterologous source [12] |
| Isoprenol (Isopentenol) | C5 alcohol substrate for the IUP. | Commercial chemical supplier |
| pSEVA228-pro4IUPi / pTET-IUPi | Plasmid vectors for constitutive (Ppro4) or inducible (PTET) IUP expression. | [12] |
| M9 Minimal Media | Defined medium for growth assays and selection. | Standard formulation |
| CRISPR-Cas9 System | For targeted knockout of the ispG gene in the MEP pathway. | [12] |
Step-by-Step Workflow
Figure 2. Workflow for IUP Functional Assay.
Generate MEP-Knockout Strain (KO1):
- Use a CRISPR-Cas9 system to disrupt the ispG gene in E. coli, which encodes (E)-4-hydroxy-3-methyl-but-2-enyl pyrophosphate (HMBPP) synthase, an essential enzyme in the native MEP pathway [12].
- Control: Validate that the KO1 strain is non-viable in M9 minimal media but viable when supplemented with 1 mM mevalonate (which feeds into the lower MVA pathway).
Clone the IUP Operon:
- Assemble a synthetic operon containing genes for ScCK, AtIPK, and IDI.
- Clone this operon into two distinct expression vectors: a constitutive plasmid (e.g., pSEVA228-pro4IUPi) and a strong inducible plasmid (e.g., pTET-IUPi) [12].
Transform and Create Engineered Strains:
- Transform the IUP plasmids into the KO1 strain to create rescue strains KO2 (constitutive) and KO3 (inducible).
Functional Rescue Assay:
- Inoculate KO2 and KO3 strains into M9 minimal media supplemented with 0.6 mM isoprenol. Do not add mevalonate.
- Incubate at 37°C with shaking and monitor optical density (OD600) over 48-72 hours.
- Expected Outcome: Successful rescue strains will exhibit growth after a lag period (e.g., ~36 hours), confirming that the IUP can solely provide essential isoprenoids for cell survival [12].
Quantitative Analysis:
- Compare the exponential growth rates of the rescued strains to wild-type E. coli.
- For quantification of isoprenoid output, couple the IUP to a lycopene production module and extract lycopene for quantification via UV/Vis spectroscopy [12].
Protocol 2: Coupling the IUP to C1-Derived Isopentenol for Lycopene Production
This protocol describes the implementation of the IUP in a wild-type E. coli host for high-yield production of a model isoprenoid, lycopene, using exogenously supplied isopentenol.
Research Reagent Solutions
| Reagent / Material | Function / Description | Source / Example |
|---|---|---|
| Lycopene Production Plasmid | Contains genes for condensation of GGPP to lycopene (CrtE, CrtI, CrtB). | Commercial repository or literature |
| Isoprenol / Prenol | Pathway substrate. Can be sourced from C1-utilizing engineered microbes. | Chemical supplier or bioreactor harvest |
| Terrific Broth (TB) / LB Media | Rich media for high-density cultivation and protein expression. | Standard formulation |
| Anhydrotetracycline (aTC) | Inducer for the PTET promoter on the pTETIUP plasmid. | Commercial supplier |
Step-by-Step Workflow
Strain Preparation:
- Co-transform wild-type E. coli (e.g., DH5α) with the pTETIUP plasmid and a lycopene production plasmid.
Culture Conditions and Induction:
- Grow the engineered strain in TB media at 37°C to mid-exponential phase (OD600 ~0.6).
- Induce IUP expression by adding a final concentration of 200 ng/mL anhydrotetracycline (aTC).
- Simultaneously, add isoprenol to a final concentration of 0.6-1.0 mM.
- Continue incubation for 24-48 hours post-induction.
Product Quantification - Lycopene Extraction and Measurement:
- Harvest cells by centrifugation.
- Extract lycopene by resuspending the cell pellet in acetone and incubating at 55°C for 15 minutes, protected from light.
- Centrifuge to remove cell debris, and transfer the acetone supernatant containing lycopene to a fresh tube.
- Measure the absorbance of the extract at 472 nm using a UV/Vis spectrophotometer.
- Calculate the lycopene concentration using the Beer-Lambert law with an extinction coefficient (ε472) of 1850 mM-1cm-1 (or as determined by a standard curve).
The Scientist's Toolkit
Table 2: Essential Research Reagents for Isoprenoid Pathway Engineering
| Category | Item | Specific Example | Critical Function |
|---|---|---|---|
| Pathway Enzymes | Choline Kinase | S. cerevisiae ScCK [12] | Phosphorylates isoprenol/prenol to monophosphate |
| Isopentenyl Phosphate Kinase | A. thaliana AtIPK [12] | Phosphorylates monophosphate to active diphosphate (IPP/DMAPP) | |
| 1-Deoxy-D-Xylulose 5-Phosphate Synthase (DXS) | P. tricornutum PtDXS [10] [39] | Rate-limiting enzyme in the native MEP pathway | |
| Genetic Tools | Constitutive / Inducible Vectors | pSEVA228 (constitutive), pTET (inducible) [12] | Controlled expression of heterologous pathways |
| CRISPR-Cas9 System | For ispG knockout [12] | Targeted gene disruption for metabolic engineering | |
| Analytical Standards | Isopentenyl Pyrophosphate (IPP) | Commercial standard | HPLC/MS quantification of pathway precursors |
| Lycopene | Commercial standard (e.g., from tomato) | Quantification of isoprenoid output via UV/Vis | |
| Specialized Substrates | Isoprenol (Isopentenol) | 3-Methyl-3-buten-1-ol | Direct substrate for the IUP |
| C1 Carbon Sources | Methanol, Formate | Feedstock for sustainable precursor production |
Integration with Broader Thesis Research
The protocols outlined herein provide a practical foundation for a broader thesis exploring the frontiers of isoprenoid pathway engineering. The IUP serves as a modular chassis that can be integrated with upstream and downstream modules:
- Upstream Integration: A primary research direction is coupling the IUP to strains engineered for isopentenol synthesis from C1 substrates. This involves introducing C1 assimilation pathways (e.g., methanol oxidation, the Calvin cycle, or synthetic formats) and engineering the host to convert central metabolites to isopentenol, creating a seamless pipeline from waste carbon to high-value isoprenoids.
- Downstream Expansion: The high-flux generation of IPP/DMAPP via the IUP can be channeled into a diverse array of terpenoid skeletons by expressing heterologous terpene synthases and cytochrome P450s [10] [12]. This allows for the production of complex pharmaceutical compounds like taxadiene (a taxol precursor) or artemisinic acid.
- Host Diversification: While demonstrated in E. coli, the IUP concept is portable. A key research avenue is its implementation in microalgae like Phaeodactylum tricornutum, which natively possess the MEP pathway and can fix CO2 [10] [39]. Engineering microalgae to utilize the IUP with wastewater-derived carbon could unlock ultra-sustainable production paradigms.
Troubleshooting and Optimization Guide
| Problem | Potential Cause | Suggested Solution |
|---|---|---|
| No growth in rescue assay (Protocol 1.4) | IUP expression too low; inefficient enzyme kinetics. | Use the stronger inducible pTETIUP plasmid. Optimize codon usage for heterologous genes. Increase isoprenol concentration to 1.0 mM. |
| Low lycopene yield (Protocol 4.2) | Imbalanced IPP/DMAPP ratio; substrate toxicity; low ATP. | Ensure IDI is co-expressed. Titrate isoprenol feeding rate to avoid toxicity. Use rich media (TB) to ensure ample ATP supply. |
| High background in non-induced cultures | Promoter leakiness in expression system. | Use tighter regulatory systems (e.g., L-rhamnose inducible). Isolate and use a clone with low basal expression. |
The Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and CRISPR-associated (Cas) systems have revolutionized genetic engineering, providing unprecedented precision and programmability for microbial metabolic engineering. Derived from bacterial adaptive immune systems, these technologies enable researchers to perform targeted genome editing, transcriptional regulation, and metabolic pathway optimization with high efficiency [40] [41]. For metabolic engineers focused on isoprenoid biosynthetic pathway optimization, CRISPR/Cas systems offer powerful tools to overcome long-standing challenges in pathway regulation, flux balancing, and host strain development [42] [43].
Isoprenoids, also known as terpenoids, represent one of the largest families of natural products with over 50,000 identified compounds possessing diverse applications in pharmaceuticals, nutraceuticals, flavors, fragrances, and advanced biofuels [44] [12] [45]. Despite their commercial significance, the industrial production of isoprenoids faces substantial challenges due to the structural complexity of these compounds, their typically low abundance in native sources, and the intricate regulation of their biosynthetic pathways in host organisms [10] [45]. The advent of CRISPR/Cas technology has accelerated engineering efforts in model microbial hosts such as Escherichia coli and Saccharomyces cerevisiae, enabling systematic optimization of both native and heterologous isoprenoid pathways through precise genetic modifications [43] [46] [41].
This application note provides detailed methodologies and protocols for implementing CRISPR/Cas systems in isoprenoid pathway engineering, including experimental workflows, reagent specifications, and representative case studies with quantitative outcomes.
CRISPR/Cas Toolkit Components for Pathway Engineering
Core Editing Systems
The foundational CRISPR/Cas toolkit comprises several specialized components that can be customized for specific metabolic engineering applications:
Cas Protein Variants: The CRISPR system centers on Cas effector proteins, with Streptococcus pyogenes Cas9 (SpCas9) serving as the most widely adopted nuclease. For applications requiring different protospacer adjacent motif (PAM) specificities or smaller protein sizes, alternatives such as Cas12a (Cpf1) and engineered compact variants like CasMINI offer distinct advantages [42]. High-fidelity versions (e.g., SpCas9-HF1, eSpCas9) incorporate mutations that reduce non-specific DNA binding, minimizing off-target effects in metabolic engineering applications [42].
Guide RNA (gRNA) Design: The single-guide RNA (sgRNA) consists of a CRISPR RNA (crRNA) component that confers target specificity through 20-nucleotide base pairing with the DNA target, fused to a trans-activating crRNA (tracrRNA) scaffold that facilitates Cas protein binding [40] [41]. For effective targeting, sgRNAs should be designed to bind non-template DNA strands at positions proximal to the start codon of metabolic genes, with careful screening to minimize off-target effects [43].
Delivery Systems: Efficient intracellular delivery of CRISPR components remains crucial for successful genome editing. Established methods include electroporation, PEG-mediated transformation, and particle bombardment (biolistics), with emerging approaches utilizing engineered viral vectors or nanoparticle complexes for enhanced efficiency [42].
Advanced CRISPR Derivatives
Beyond basic gene editing, advanced CRISPR derivatives enable sophisticated metabolic pathway control:
CRISPR Interference (CRISPRi): Utilizing a catalytically deactivated Cas9 (dCas9), CRISPRi blocks transcription by sterically hindering RNA polymerase, enabling tunable gene knockdown without permanent DNA modification [43]. This approach is particularly valuable for transiently repressing competitive metabolic pathways that would be lethal if permanently inactivated.
CRISPR Activation (CRISPRa): By fusing dCas9 to transcriptional activation domains (e.g., VP64, p65AD), CRISPRa enhances gene expression, facilitating overexpression of rate-limiting enzymes in isoprenoid biosynthetic pathways without the need for promoter replacements [42].
Base and Prime Editing: These DSB-free editing systems enable precise nucleotide conversions (base editors) or small insertions/deletions (prime editors), allowing for fine-tuning of enzyme activity and regulatory elements without introducing double-strand breaks that can cause cytotoxicity [42].
Application Protocols for Isoprenoid Pathway Engineering
Protocol 1: CRISPRi-Mediated Multiplex Regulation of the MVA Pathway inE. coli
This protocol details the implementation of CRISPRi for multiplexed downregulation of endogenous genes to enhance flux through the heterologous mevalonate (MVA) pathway for isoprenol production in E. coli [43].
Experimental Workflow
Materials and Reagents
- Bacterial Strains: E. coli strain heterologously expressing either the native MVA pathway or IPP-bypass pathway for isoprenol production [43]
- Plasmids: pdCas9-Marraffini (JBEI-18706) encoding dCas9 under Ptet inducible promoter [43]
- gRNA Expression Vectors: Golden Gate-compatible vectors with constitutive promoters for sgRNA expression
- Culture Media: M9 minimal medium with 2% glucose; appropriate antibiotics for selection
- Induction Reagents: Anhydrotetracycline (aTc) for dCas9 induction
- Analytical Equipment: GC-MS system for isoprenol quantification
Step-by-Step Procedure
sgRNA Design and Selection:
- Identify target genes competing for precursors (e.g., dxs, idi, ispD, ispF, ispG, ispH) and genes in central metabolism (e.g., pfkA, pfkB, pykA, pykF)
- Design 30-bp sgRNA sequences using Benchling with the following criteria:
- Target the non-template DNA strand
- Position proximal to start codon or promoter regions
- Minimize off-target potential through comprehensive genome scanning
- Select 5-10 top sgRNAs for initial screening
Plasmid Construction:
- Assemble sgRNA expression cassettes using Golden Gate assembly with BsaI restriction sites
- Clone individual sgRNAs into the pdCas9-Marraffini backbone
- Verify constructs by Sanger sequencing before transformation
Strain Transformation and Screening:
- Transform individual CRISPRi plasmids into production E. coli strains via heat shock or electroporation
- Plate transformed cells on LB agar with appropriate antibiotics and incubate at 30°C overnight
- Pick 3-5 colonies per transformation for small-scale screening in 96-deep well plates
- Culture in M9 minimal medium with 2% glucose and induce with 100 ng/mL aTc at OD600 = 0.4-0.6
- Incubate for 48-72 hours at 30°C with shaking at 250 rpm
Isoprenol Quantification:
- Harvest 1 mL culture samples and centrifuge at 13,000 × g for 5 minutes
- Extract isoprenol from supernatant with ethyl acetate (1:1 v/v)
- Analyze extracts by GC-MS using standard calibration curves
- Normalize titers to OD600 and identify top-performing sgRNAs
Multiplexed Strain Construction:
- Assemble a multiplex gRNA array containing 3-5 top-performing sgRNAs
- Transform the multiplex CRISPRi construct into production strains
- Validate multiplex repression by RT-qPCR of target genes
Bioreactor Scale-Up:
- Transfer top-performing strains to 2-L bioreactors with M9 minimal medium
- Maintain dissolved oxygen at 30%, pH at 7.0, and temperature at 30°C
- Induce with 100 ng/mL aTc at mid-exponential phase
- Implement fed-batch feeding with concentrated glucose solution to maintain ~2 g/L
- Monitor isoprenol production over 72-96 hours
Expected Results and Troubleshooting
- Expected Outcomes: Single sgRNA strains should show 1.5-3 fold titer improvements; multiplexed strains typically achieve 3-4.5 fold enhancements [43]
- Troubleshooting:
- Low Editing Efficiency: Optimize aTc concentration (50-200 ng/mL), verify dCas9 expression by Western blot
- Poor Growth: Titrate sgRNA expression, check for essential gene repression
- Variable Titers: Maintain consistent induction OD600, standardize extraction protocols
Protocol 2: CRISPR-Cas9 Mediated Genome Integration of Carotenoid Pathways
This protocol describes CRISPR-Cas9 mediated multiplex integration of the β-carotene biosynthetic pathway into the E. coli genome, enabling stable carotenoid production without selection pressure [46].
Experimental Workflow
Materials and Reagents
- Bacterial Strains: E. coli MG1655 ΔbioA::λ-Red (EcPHE) for recombineering [46]
- Plasmids: pCas9cur (Cas9 expression, λ-Red, temperature-sensitive origin) [46]
- Pathway Templates: Donor DNA containing crtE, crtI, crtYB genes with homology arms
- Culture Media: LB medium; appropriate antibiotics; L-arabinose for λ-Red induction
- Analytical Equipment: HPLC system with photodiode array detector, spectrophotometer
Step-by-Step Procedure
Donor DNA and gRNA Design:
- Design 500-1000 bp homology arms flanking the integration site (e.g., galK, yghB loci)
- Assemble β-carotene pathway genes (crtE, crtI, crtYB) with optimized promoters and RBS
- Design gRNAs targeting neutral genomic sites with minimal off-target potential
Editing Plasmid Assembly:
- Clone gRNA expression cassette into pCas9cur derivative
- Prepare linear donor DNA fragment by PCR or synthesis
- Co-transform gRNA plasmid and donor DNA into EcPHE expressing λ-Red proteins
Cas9-induced DSB and HDR:
- Culture transformed cells at 30°C with appropriate antibiotics to OD600 = 0.3-0.5
- Induce λ-Red recombineering with 10 mM L-arabinose for 1 hour
- Induce Cas9 expression with 0.2 mM IPTG for 2 hours
- Plate cells on selective media and incubate at 30°C for 36-48 hours
Plasmid Curing:
- Streak colonies on non-selective media and incubate at 37°C overnight
- Screen for antibiotic-sensitive clones to confirm plasmid loss
- Verify pathway integration by colony PCR and sequencing
Combinatorial Pathway Modulation:
- Implement iterative CRISPR editing to modulate MEP pathway genes (dxs, idi, ispDF)
- Target central metabolic genes (pgi, zwf, gnd) to enhance precursor supply
- Use ssDNA donors for promoter replacements and RBS optimization
Carotenoid Extraction and Analysis:
- Harvest cells by centrifugation and extract pigments with acetone:methanol (7:3 v/v)
- Analyze extracts by HPLC with C30 column and compare to authentic standards
- Quantify β-carotene concentration using extinction coefficient (E1%1cm = 2592 at 450 nm)
Expected Results and Troubleshooting
- Expected Outcomes: Initial integration should yield 2-5 mg/L β-carotene; combinatorial optimization can increase titers to >20 mg/L [46]
- Troubleshooting:
- Low Integration Efficiency: Optimize homology arm length, verify λ-Red expression
- Cytotoxicity: Titrate Cas9 expression, use high-fidelity Cas9 variants
- Uneven Pathway Expression: Fine-tune with RBS libraries, promoter engineering
Research Reagent Solutions for Isoprenoid Engineering
Table 1: Essential Research Reagents for CRISPR-mediated Isoprenoid Pathway Engineering
| Reagent Category | Specific Examples | Function | Application Notes |
|---|---|---|---|
| Cas Effectors | SpCas9, FnCas12a, CasMINI, dCas9 | DNA cleavage or binding | Cas12a preferred for AT-rich regions; dCas9 for CRISPRi/a [42] |
| gRNA Expression Systems | U6 promoters, tRNA scaffolds | Guide RNA expression | tRNA-sgRNA systems enhance processing in bacteria [41] |
| Delivery Vectors | pdCas9-Marraffini, pCas9cur | Component delivery | Temperature-sensitive origins enable plasmid curing [43] [46] |
| Donor DNA Templates | dsDNA with homology arms, ssDNA oligos | Homology-directed repair | 500-1000 bp arms for dsDNA; 90 nt for ssDNA in E. coli [46] |
| Induction Systems | aTc-inducible Ptet, L-arabinose inducible | Controlled expression | Fine-tune expression timing to balance editing and viability [43] |
| Pathway Modules | MVA pathway genes, IUP enzymes, carotenoid clusters | Isoprenoid production | IPP-bypass pathway reduces toxicity [12] [43] |
| Analytical Tools | GC-MS, HPLC-PDA, RT-qPCR | Product quantification & validation | RT-qPCR essential for verifying CRISPRi repression [43] |
Case Studies and Quantitative Outcomes
Case Study 1: Isoprenol Production via CRISPRi-tuned IPP-bypass Pathway
Table 2: Quantitative Outcomes of CRISPRi-enhanced Isoprenol Production in E. coli [43]
| Engineering Strategy | Host Strain | Target Genes | Isoprenol Titer (g/L) | Fold Improvement |
|---|---|---|---|---|
| Baseline Strain | MVA Pathway E. coli | None | 0.40 ± 0.05 | 1.0 |
| Single sgRNA | MVA Pathway E. coli | dxs | 0.85 ± 0.08 | 2.1 |
| Multiplexed sgRNA | MVA Pathway E. coli | dxs, idi, ispDF | 1.45 ± 0.12 | 3.6 |
| Baseline IPP-bypass | IPP-bypass E. coli | None | 0.53 ± 0.06 | 1.0 |
| Optimized CRISPRi | IPP-bypass E. coli | glnA, ldhA, pfkA | 1.82 ± 0.19 | 3.4 |
| Fed-batch Scale-up | IPP-bypass + Multiplex CRISPRi | glnA, ldhA, pfkA | 12.4 ± 1.3 | 23.4 |
Case Study 2: β-Carotene Production via Combinatorial CRISPR Engineering
Implementation of CRISPR-Cas9 mediated genome editing for β-carotene pathway integration and optimization resulted in significant production enhancements [46]:
- Single-step integration of the complete β-carotene pathway (crtE, crtI, crtYB) achieved initial titers of 3.2 mg/L
- Combinatorial modulation of the MEP pathway through dxs overexpression and ispF repression increased titers to 18.7 mg/L
- Additional engineering of central metabolic genes (pgi knockout to enhance PPP flux) further improved production to 25.6 mg/L
- The complete process required only 3 iterative editing cycles over 6 days, demonstrating the accelerated strain development timeline enabled by CRISPR-Cas9 systems
CRISPR/Cas systems have emerged as indispensable tools for isoprenoid pathway engineering, enabling precise genome editing, multiplex gene regulation, and rapid strain optimization. The protocols outlined in this application note provide researchers with robust methodologies for implementing these advanced genetic toolkits in microbial hosts. As CRISPR technology continues to evolve, emerging capabilities such as base editing, prime editing, and CRISPR-mediated biosensing will further expand the metabolic engineer's toolbox, enabling more sophisticated control over isoprenoid biosynthetic pathways and accelerating the development of microbial cell factories for sustainable production of high-value isoprenoid compounds [42] [41].
Within the broader context of isoprenoid biosynthetic pathway engineering, the successful production of artemisinin stands as a seminal case study. Artemisinin, a potent antimalarial sesquiterpene lactone, is derived from the plant Artemisia annua L [47] [48]. Its complex structure, featuring an endoperoxide bridge, makes chemical synthesis economically unfeasible, thereby positioning metabolic engineering as the primary strategy for achieving a stable, scalable supply [49] [50]. This application note details the foundational biosynthetic pathways, presents a successful heterologous production case study in the moss Physcomitrella patens, and provides associated experimental protocols and reagent solutions for researchers in the field.
Biosynthetic Pathway of Artemisinin
Artemisinin biosynthesis occurs primarily in the glandular trichomes of Artemisia annua and originates from universal isoprenoid precursors [47] [49]. The pathway demonstrates a key feature in plant metabolism: the potential collaboration between two distinct precursor pathways, the cytosolic Mevalonate (MVA) and the plastidial Methylerythritol Phosphate (MEP) pathways, both supplying Isopentenyl pyrophosphate (IPP) for sesquiterpene biosynthesis [47] [49].
The committed pathway to artemisinin involves several enzymatic steps, culminating in a non-enzymatic photo-oxidation that forms the crucial endoperoxide bridge [49].
Diagram: The Artemisinin Biosynthetic Pathway. The pathway initiates from universal isoprenoid precursors supplied by the MVA or MEP pathways. Key enzymatic steps are catalyzed by ADS, CYP71AV1, and DBR2, culminating in a non-enzymatic photo-oxidation to form artemisinin [47] [49].
Case Study: Stable Production inPhyscomitrella patens
The fluctuating supply and price of artemisinin from A. annua cultivation created an urgent need for a stable, scalable production platform [50]. The moss Physcomitrella patens was selected as a photosynthetic chassis due to its haploid genome, high efficiency of homologous recombination, and capability for scalable bioreactor cultivation [50]. This case study demonstrates the successful metabolic engineering of the entire dihydroartemisinic acid (DHAA) pathway in a heterologous plant system.
Quantitative Production Metrics
Table 1: Key Production Metrics from the P. patens Case Study [50]
| Parameter | Result | Context & Significance |
|---|---|---|
| Artemisinin Titer | 0.21 mg/g Dry Weight (DW) | Measured in transgenic moss lines; a significant initial production level. |
| Production Time | 3 days | Rapid production after a short cultivation period. |
| Key Pathway Product | Dihydroartemisinic Acid (DHAA) | The final enzymatic precursor, which converts non-enzymatically to artemisinin. |
| Transformation System | Direct in vivo DNA assembly | Enabled simultaneous integration of five biosynthetic genes into the genome. |
Detailed Experimental Protocol
This section outlines the core protocol for engineering artemisinin production in P. patens, from vector construction to metabolite analysis.
Protocol Workflow
Diagram: Experimental Workflow for Artemisinin Production in P. patens. The process involves constructing the biosynthetic pathway, transforming moss protoplasts, and validating transgenic lines [50].
Step-by-Step Procedure
Step 1: Pathway Design and Gene Assembly
- Construct Design: Assemble a multi-gene construct containing the five key enzymes for DHAA biosynthesis:
- Amorphadiene Synthase (ADS)
- Cytochrome P450 CYP71AV1
- Alcohol Dehydrogenase 1 (ADH1)
- Double-bond Reductase 2 (DBR2)
- Aldehyde Dehydrogenase 1 (ALDH1) [50].
- Genetic Optimization: Codon-optimize the genes (particularly CYP71AV1, ADH1, DBR2, and ALDH1) for P. patens codon usage [50].
- Vector Assembly: Use a transformation system that allows for the direct in vivo assembly of multiple DNA fragments via homologous recombination. Flank the construct with genomic sequences from the Pp108 locus for targeted integration [50].
Step 2: Moss Protoplast Preparation
- Culture Conditions: Grow P. patens (Gransden ecotype) on solid or liquid PhyB media at 25°C under continuous light [50].
- Protoplast Isolation:
- Harvest approximately 1.5 g of 5-day-old moss tissue.
- Digest cell walls using 0.5% Driselase enzyme solution in 8.5% mannitol. Use 1 mL solution per 40 mg tissue.
- Incubate at room temperature for 30-60 min with gentle shaking.
- Filter the digest through a 100-µm mesh and collect protoplasts by centrifugation at 150-200 × g for 4 minutes.
- Wash the pellet twice with protoplast wash solution (8.5% mannitol, 10 mM CaCl₂) [50].
- Suspension: Resuspend protoplasts in MMM solution (9.1% d-mannitol, 10% MES, 15 mM MgCl₂) at a density of 1.6 × 10⁶ protoplasts/mL [50].
Step 3: Genetic Transformation
- Transformation Mix: Combine 300 µL of protoplast suspension with 10 µg of total DNA and 300 µL of PEG solution.
- Heat Shock: Incubate the mixture for 5 minutes at 45°C, followed by 5 minutes at room temperature.
- Dilution and Plating: Gradually dilute the transformation mix with 8.5% d-mannitol. Plate the protoplasts on protoplast regeneration media (PRMB) overlaid with cellophane. Incubate at 25°C under continuous light for 5-7 days [50].
Step 4: Selection and Regeneration
- Antibiotic Selection: After initial regeneration, transfer the protoplasts to PhyB media containing the appropriate antibiotic (e.g., G418 or hygromycin) for selection of transformants.
- Stable Line Isolation: Cycle between selection and non-selection media with biweekly subculturing, blending the tissue in sterile water between cycles, until stable transgenic lines are established [50].
Step 5: Molecular Validation
- PCR Analysis: Confirm the successful integration of the transgenes into the P. patens genome using genomic PCR with gene-specific primers.
- Expression Check: Verify the expression of the introduced pathway genes via RT-PCR or quantitative RT-PCR [50].
Step 6: Metabolite Analysis
- Extraction: Harvest transgenic moss tissue and extract metabolites using suitable organic solvents (e.g., hexane or methanol).
- Quantification: Analyze artemisinin and precursor content (e.g., DHAA) using techniques such as High-Performance Liquid Chromatography (HPLC) or Liquid Chromatography-Mass Spectrometry (LC-MS). The reported production can reach 0.21 mg/g DW of artemisinin [50].
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Research Reagents for Artemisinin Pathway Engineering
| Reagent / Solution | Function / Application | Example / Specification |
|---|---|---|
| PhyB Media | Standard growth medium for P. patens cultivation. | Contains Ca(NO₃)₂, MgSO₄·7H₂O, KH₂PO₄ buffer (pH 6.5), and trace elements [50]. |
| Driselase Enzyme | Digests cell wall to produce protoplasts for transformation. | 0.5% solution in 8.5% mannitol [50]. |
| Protoplast Wash Solution | Washes and maintains protoplast integrity post-digestion. | 8.5% mannitol, 10 mM CaCl₂ [50]. |
| MMM Solution | Resuspension solution for protoplasts prior to transformation. | 9.1% d-mannitol, 10% MES, 15 mM MgCl₂ [50]. |
| PEG Solution | Facilitates DNA uptake into protoplast membranes. | Used in equal volume to protoplast suspension for transformation [50]. |
| Protoplast Regeneration Media (PRMT/PRMB) | Supports cell wall reformation and growth of transformed protoplasts. | Two-layer system (top and bottom) for optimal recovery [50]. |
| Antibiotic Selection Markers | Selects for successfully transformed lines. | G418 or hygromycin resistance genes driven by constitutive promoters [50]. |
| Codon-Optimized Genes | Ensures high expression of heterologous enzymes in the host. | Genes like CYP71AV1, DBR2 optimized for P. patens codon usage [50]. |
The successful engineering of artemisinin biosynthesis in Physcomitrella patens provides a validated and scalable protocol for the sustainable production of this critical pharmaceutical. This case study exemplifies the power of metabolic engineering applied to isoprenoid pathways, demonstrating that a heterologous photosynthetic system can be engineered to produce complex secondary metabolites. The detailed methodologies and reagent solutions outlined herein serve as a practical framework for researchers aiming to replicate or build upon this work for the production of artemisinin or other high-value isoprenoids.
Navigating Bottlenecks: Troubleshooting and Flux Optimization
Identifying and Alleviating Metabolic Bottlenecks and Cytotoxicity
Within the framework of isoprenoid biosynthetic pathway engineering, a significant challenge is the inherent inefficiency of converting central metabolic precursors into the universal isoprenoid building blocks, isopentenyl diphosphate (IPP) and dimethylallyl diphosphate (DMAPP). This inefficiency often manifests as metabolic bottlenecks, where the suboptimal activity of a specific enzyme restricts carbon flux through the entire pathway. Furthermore, the accumulation of certain pathway intermediates or the final isoprenoid products can exert cytotoxicity, inhibiting cell growth and limiting overall production titers [51]. This Application Note details the identification of common bottlenecks in engineered isoprenoid pathways and provides validated, detailed protocols to alleviate these constraints, thereby enhancing the performance of microbial cell factories.
Identifying Common Metabolic Bottlenecks
A critical first step in pathway optimization is pinpointing the specific enzymatic steps that limit flux. The following table summarizes common bottlenecks, their indicators, and standard identification methods.
Table 1: Common Metabolic Bottlenecks in Engineered Isoprenoid Pathways
| Bottleneck Category | Specific Example | Key Indicators | Common Identification Methods |
|---|---|---|---|
| Precursor Supply | Acetyl-CoA/IPP/DMAPP availability | Accumulation of upstream precursors (e.g., acetyl-CoA), low downstream product titers [52] | Metabolite profiling (LC-MS), 13C Metabolic Flux Analysis [34] [52] |
| Rate-Limited Enzyme Activity | Mevalonate kinase (MK), HMG-CoA reductase (HMGR) | Accumulation of the enzyme's substrate (e.g., mevalonate, HMG-CoA) [52] | Targeted proteomics, enzyme activity assays, metabolite profiling [34] [52] |
| Inadequate Cofactor Supply | NADPH/ATP availability | Imbalanced NADPH/NADP+ ratio, reduced pathway efficiency [34] | Cofactor profiling, overexpression of cofactor regeneration enzymes [34] |
| Competitive Pathway Drain | Native sterol biosynthesis consuming FPP | Decreased yield of target non-sterol isoprenoid despite high precursor flux [51] | Transcriptomics, knock-out/knock-down of competing genes [34] [51] |
The logical workflow for identifying these bottlenecks begins with cultivating the engineered production strain and systematically analyzing samples to pinpoint the limiting step, as outlined below.
Experimental Protocol: Metabolite Profiling for Bottleneck Identification
This protocol is adapted from methods used to identify IPP accumulation in E. coli [52].
Key Materials:
- Engineered microbial production strain (e.g., E. coli KG1 [52])
- Appropriate culture medium (e.g., M9 or LB with selective antibiotics)
- Quenching solution (e.g., 60:40 v/v methanol:water at -40 °C)
- Extraction solvent (e.g., cold methanol or acetonitrile)
- Internal standards (e.g., deuterated mevalonate, 13C-labeled IPP)
- LC-MS/MS system (e.g., UHPLC coupled to a triple quadrupole mass spectrometer)
Procedure:
- Culture and Sampling: Inoculate the production strain in triplicate and cultivate under optimal conditions. At defined time points (e.g., 0, 6, 12, 24, 48 hours), aseptically withdraw a precise volume of culture (e.g., 1 mL).
- Rapid Quenching: Immediately transfer the sample to a pre-chilled tube containing 4 mL of quenching solution to halt metabolism. Incubate on ice for 10 minutes.
- Cell Pellet Collection: Centrifuge the quenched sample at high speed (e.g., 10,000 × g, 5 min, -4 °C). Carefully discard the supernatant.
- Metabolite Extraction: Resuspend the cell pellet in 1 mL of cold extraction solvent. Vortex vigorously for 1 minute. Incubate at -20 °C for 1 hour, then centrifuge (14,000 × g, 10 min, 4 °C).
- Sample Analysis: Transfer the supernatant to a new vial. Analyze the extracted metabolites using a targeted LC-MS/MS method optimized for the separation and detection of MVA/MEP pathway intermediates (e.g., mevalonate, mevalonate-5-phosphate, IPP, DMAPP) based on their known mass transitions and retention times. Use internal standards for quantification.
Data Interpretation: A significant accumulation of a specific intermediate (e.g., an 8-fold higher concentration of IPP compared to other intermediates [52]) strongly indicates that the subsequent enzymatic step (e.g., dephosphorylation of IPP by NudB) is a primary bottleneck.
Strategies and Protocols for Alleviating Bottlenecks and Cytotoxicity
Once a bottleneck is identified, targeted strategies can be implemented. The following table compares the core approaches.
Table 2: Strategies for Alleviating Bottlenecks and Cytotoxicity
| Strategy | Core Principle | Key Example | Reported Efficacy |
|---|---|---|---|
| Enzyme & Pathway Engineering | Overexpress rate-limiting enzymes; engineer synthetic pathways [34] [51] | Optimizing NudB Shine-Dalgarno sequence to enhance IPP conversion [52] | 60% increase in 3-methyl-3-buten-1-ol yield; 9-fold increase in NudB protein [52] |
| Membrane & Compartment Engineering | Increase storage capacity for hydrophobic products; compartmentalize pathways [51] | Knockdown of cell division genes to increase total membrane area in E. coli [51] | Enhanced production of carotenoids and violacein derivatives [51] |
| Salvage & Orthogonal Pathways | Utilize simpler, decoupled pathways for precursor synthesis [53] [54] | Introducing Isopentenol Utilization Pathway (IUP) in C. reinhardtii [54] | 8.6-fold increase in IPP; 23-fold increase in limonene production [54] |
The relationship between these strategies and their impact on the engineered isoprenoid pathway is visualized below.
Experimental Protocol: Implementing an Orthogonal Salvage Pathway
The Isopentenol Utilization Pathway (IUP) is a two-step salvage pathway that efficiently converts exogenous isoprenol and prenol into IPP and DMAPP, bypassing complex native regulation [53] [54].
Key Materials:
- Chlamydomonas reinhardtii strain (or other host of choice)
- Codon-optimized genes for ScCK (alcohol kinase) and AtIPK (isopentenyl phosphate kinase) [54]
- Chloroplast transformation vectors for C. reinhardtii
- opt2 culture medium
- Isoprenol (prenol)
- Dodecane (for in situ extraction of limonene, if applicable)
Procedure:
- Construct Assembly: Clone the ScCK and AtIPK genes into an appropriate chloroplast expression vector for C. reinhardtii. Constructs can include fluorescent protein fusions (e.g., YFP, CFP) to facilitate screening of high-expression transformants [54].
- Strain Transformation: Introduce the expression construct into the C. reinhardtii chloroplast via biolistic or glass bead transformation. Select transformants on spectinomycin-containing plates.
- Screening: Screen for positive transformants using fluorescence microscopy or flow cytometry to identify clones with strong fluorescence, indicating high expression of the IUP enzymes.
- Fed-Batch Cultivation: Inoculate the transgenic algal strain in opt2 medium. Supplement the culture with a fed-batch of isoprenol (e.g., 10 mM final concentration) to provide the pathway substrate [54].
- Product Analysis: Monitor cell growth. For volatile terpenoids like limonene, use a dodecane overlay during cultivation to capture the product. Analyze the dodecane phase and cell pellets using gas chromatography (GC-MS or GC-FID) to quantify terpenoid production.
Data Interpretation: Successful implementation of the IUP is indicated by a significant increase in total IPP/DMAPP titers and a consequent boost in the production of the target terpenoid (e.g., limonene) compared to the control strain without the IUP [54].
The Scientist's Toolkit: Key Research Reagent Solutions
The following table lists essential reagents and tools for implementing the protocols described in this note.
Table 3: Research Reagent Solutions for Isoprenoid Pathway Engineering
| Reagent/Tool | Function/Application | Example Source/Description |
|---|---|---|
| Fosmidomycin | Inhibitor of the native MEP pathway (DXR enzyme); used to decouple native metabolism for testing orthogonal pathways [53] | Commercial chemical inhibitor |
| NudB Phosphatase | Promiscuous phosphatase in E. coli; catalyzes the dephosphorylation of IPP/DMAPP to form isopentenol/prenol [52] | E. coli native enzyme; can be overexpressed and engineered |
| Isopentenol (Isoprenol) | Membrane-permeable precursor for the Isopentenol Utilization Pathway (IUP) [53] [54] | Commercial chemical substrate |
| ScCK & AtIPK Enzymes | Core two-enzyme system for the IUP; ScCK performs first phosphorylation, AtIPK the second [54] | Codon-optimized genes for heterologous expression |
| CRISPR-Cas9 Systems | Genome editing tool for knock-out of competing pathways or genomic integration of heterologous genes [10] [24] | Specific tools for host organism (e.g., yeast, E. coli, microalgae) |
| Strong Constitutive/Tunable Promoters | Drive high-level or regulated expression of pathway enzymes (e.g., lacUV5, trc) [52] | Plasmid-based or genomic expression systems |
Balancing Cofactor Supply and Demand for Enhanced Pathway Flux
In the context of a broader thesis on isoprenoid biosynthetic pathway engineering, the critical challenge of balancing metabolic cofactors emerges as a central research focus. Isoprenoid biosynthesis, whether through the mevalonate (MVA) pathway or methylerythritol phosphate (MEP) pathway, imposes significant demands on cellular cofactors, particularly ATP and NADPH [10] [34]. The inherent mismatch between the cofactor requirements of engineered pathways and the host's native supply networks often creates metabolic bottlenecks that severely limit flux toward target compounds. This application note details standardized protocols and analytical frameworks for diagnosing and resolving these cofactor imbalances, enabling researchers to achieve substantial yield improvements in microbial isoprenoid production platforms.
Cofactor Demands in Isoprenoid Biosynthetic Pathways
The biosynthesis of isoprenoid precursors requires distinct cofactor investment patterns depending on the pathway utilized. Understanding these requirements is essential for targeted engineering approaches.
Table 1: Cofactor Requirements for Major Isoprenoid Precursor Pathways
| Pathway | Key Cofactors | ATP Consumption per IPP | NADPH Consumption per IPP | Notable Features |
|---|---|---|---|---|
| Classical MVA Pathway | ATP, NADPH | 3 ATP | 2 NADPH | High ATP demand; canonical pathway in yeast [55] |
| Archaeal MVA Pathway | ATP, NADPH | 2 ATP | 2 NADPH | Reduced ATP requirement; potential oxygen sensitivity [5] |
| MEP Pathway | ATP, NADPH, CTP | ~2.5 ATP (equiv.) | 2 NADPH | Dependent on G3P/pyruvate; plastidic localization [10] [56] |
| Isoprenol Utilization Pathway | ATP | 2 ATP | 0 NADPH | Bypasses native regulation; requires exogenous alcohol [57] |
The diagrams below illustrate the cofactor demand points and engineering strategies for the two primary native pathways.
Quantitative Assessment of Cofactor Limitations
Systematic evaluation of cofactor limitations requires both computational and experimental approaches. The following data, compiled from recent studies, illustrates the significant production gains achievable through cofactor balancing.
Table 2: Production Enhancements from Cofactor Engineering Strategies
| Engineering Strategy | Host Organism | Target Product | Production Titer Enhancement | Key Cofactor Addressed |
|---|---|---|---|---|
| Feedback-insensitive Acs + CoA enhancement [55] | S. cerevisiae | Mevalonate | 3,830 mg/L (360-fold increase) | Acetyl-CoA/ATP |
| Archaeal MVA pathway [5] | E. coli | Lycopene, β-Farnesene | Comparable to eukaryotic pathway | ATP |
| NADPH/NADP⁺ ratio modulation [34] | E. coli | Squalene | Significant yield improvement | NADPH |
| ER9 repression + pantothenate kinase [55] | S. cerevisiae | Mevalonate | 13.3 g/L in bioreactors | Acetyl-CoA |
| Isopentenol utilization pathway [57] | S. cerevisiae | Linalool analogs | Enabled novel analog production | ATP/NADPH bypass |
Experimental Protocols for Cofactor Balancing
Protocol: Enhancing Acetyl-CoA Supply in Yeast
This protocol outlines the implementation of a PDH bypass with a feedback-insensitive acetyl-CoA synthetase to enhance acetyl-CoA supply for mevalonate production in S. cerevisiae [55].
Research Reagent Solutions:
- E. coli-S. cerevisiae shuttle plasmid with URA3 marker (pJLA121 series)
- Plasmid pVSLS22 expressing Ef-mvaE, Ef-mvaS, and Se-acsL641P
- S. cerevisiae strain with ADH1, ADH4, GPD1, and GPD2 deletions
- SC-Ura medium (3 g/L Difco yeast nitrogen base, 10 g/L ammonium sulfate, 400 μM inositol, 4 g/L amino acid mix)
- (±)-Mevalonolactone analytical standard (97%, ACROS Organics)
Procedure:
- Strain Transformation: Transform the S. cerevisiae knockout strain with plasmid pVSLS22 using standard lithium acetate transformation. Select transformants on SC-Ura plates.
- Pre-culture Preparation: Inoculate 5 mL of SC-Ura medium with a single colony and incubate at 30°C with shaking at 250 rpm for 24 hours.
- Production Culture: Dilute the pre-culture to OD600 = 0.1 in 50 mL of fresh SC-Ura medium in a 250 mL baffled flask. Incubate at 30°C with shaking at 250 rpm for 72 hours.
- Metabolite Analysis: Collect 1 mL culture samples at 24-hour intervals. Centrifuge at 13,000 × g for 5 minutes and analyze the supernatant for mevalonate content using HPLC with an Aminex HPX-87H column maintained at 50°C, using 5 mM H₂SO₄ as mobile phase at 0.6 mL/min.
Protocol: Implementing the Energy-Efficient Archaeal MVA Pathway
This protocol describes the expression of the archaeal MVA pathway from Methanosarcina mazei in E. coli to reduce ATP consumption during isoprenoid production [5].
Research Reagent Solutions:
- Plasmid pBAD-mMPcomPMDh expressing full archaeal MVA pathway genes (MM_1525, MM_1524, MM_1526, MM_1871, MM_1763, JW2857, MM_1762)
- Plasmid pACYC-IBE expressing lycopene biosynthetic genes (crtE, crtI, crtB)
- E. coli strain with engineered MEP pathway
- LB medium supplemented with appropriate antibiotics (ampicillin, chloramphenicol)
- L-Arabinose for induction
Procedure:
- Strain Construction: Co-transform E. coli host strain with plasmids pBAD-mMPcomPMDh and pACYC-IBE. Select transformants on LB agar plates containing both ampicillin (100 μg/mL) and chloramphenicol (34 μg/mL).
- Culture Conditions: Inoculate a single colony into 5 mL of LB medium with antibiotics and grow overnight at 37°C with shaking at 250 rpm. Dilute the culture to OD600 = 0.05 in fresh medium and grow to OD600 = 0.6.
- Pathway Induction: Add L-arabinose to a final concentration of 0.2% (w/v) to induce expression of the archaeal MVA pathway. Continue incubation for 48 hours at 30°C.
- Oxygen Sensitivity Testing: For evaluating oxygen sensitivity, divide cultures after induction and maintain under either aerobic (250 rpm shaking) or oxygen-limiting (static) conditions.
- Product Quantification: Harvest cells by centrifugation, extract lycopene with acetone, and measure absorbance at 472 nm. Compare production levels between strains harboring archaeal versus eukaryotic MVA pathways.
Protocol: Modulating NADPH Supply
This protocol outlines strategies to enhance NADPH availability for isoprenoid biosynthesis, particularly important for reactions catalyzed by HMG-CoA reductase and DXR [34].
Research Reagent Solutions:
- Plasmid systems for overexpression of pentose phosphate pathway enzymes (e.g., glucose-6-phosphate dehydrogenase, zwf)
- Plasmid for heterologous expression of NADH kinase (pos5)
- E. coli or S. cerevisiae production strain
- Defined medium with controlled carbon sources
Procedure:
- Strain Engineering: Introduce plasmid-based expression of zwf or pos5 genes into the isoprenoid production host using standard transformation protocols.
- Cultivation for NADPH Enhancement: Grow engineered strains in defined medium with the appropriate carbon source. For E. coli, use M9 minimal medium; for yeast, use synthetic complete medium.
- Cofactor Analysis: Harvest cells during mid-exponential phase. Quench metabolism rapidly and extract intracellular cofactors using cold acetonitrile/methanol. Analyze NADPH/NADP⁺ ratios using LC-MS or enzymatic assays.
- Flux Analysis: For 13C metabolic flux analysis, grow cells in 13C-labeled glucose and analyze labeling patterns in pathway intermediates using GC-MS or IRMS to quantify flux through NADPH-generating pathways [56].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Cofactor Engineering in Isoprenoid Pathways
| Reagent / Tool | Function / Application | Example Sources |
|---|---|---|
| Feedback-insensitive Acs (Se-acsL641P) | Enhances acetyl-CoA supply from acetate; resistant to inhibition [55] | Salmonella enterica |
| Archaeal MVA Pathway Enzymes | Reduces ATP consumption for IPP synthesis by 33% [5] | Methanosarcina mazei |
| Isopentenyl Phosphate Kinase (IPK) | Enables isoprenol utilization pathway; bypasses native regulation [57] | Arabidopsis thaliana |
| Pantothenate Kinase (CAB1) | Enhances coenzyme A biosynthesis; increases acetyl-CoA pool [55] | Saccharomyces cerevisiae |
| Bifid Shunt Enzymes (Xfp, AckA) | Provides carbon-conserving route to acetyl-CoA [55] | Bifidobacterium spp. |
| ERG9 Repressible Promoter System | Downregulates competing squalene synthesis; redirects flux [55] | pMET3 promoter system |
Integrated Workflow for Cofactor Balancing
The following diagram illustrates a systematic workflow for diagnosing and addressing cofactor limitations in engineered isoprenoid pathways.
Systems biology approaches, which integrate multiple omics technologies, are revolutionizing the diagnosis and understanding of complex biological systems. By combining transcriptomics and metabolomics, researchers can gain a holistic view of cellular processes, from gene expression to functional metabolic phenotypes. This is particularly valuable in the context of diagnosing perturbations in intricate biosynthetic pathways, such as the isoprenoid biosynthetic pathway. Isoprenoids, a vast class of natural products with significant pharmaceutical and industrial applications, are synthesized via two major pathways: the mevalonate (MVA) pathway and the methylerythritol phosphate (MEP) pathway [10] [4]. Disruptions in these pathways can indicate specific disease states or successful genetic engineering outcomes. This application note details protocols for using integrated transcriptomics and metabolomics to diagnose and analyze the status of the isoprenoid biosynthetic pathway, providing a framework for researchers and drug development professionals.
Experimental Workflows and Protocols
Integrated Omics Workflow for Pathway Diagnosis
The following diagram illustrates the comprehensive workflow for diagnosing isoprenoid pathway activity using transcriptomics and metabolomics.
Detailed Experimental Protocols
Metabolomics Sample Preparation and LC-FTMS Analysis
Objective: To obtain a comprehensive, untargeted profile of metabolites, including intermediates of the MVA and MEP pathways.
Materials:
- Growth Media: Minimal medium with defined sulfur sources (e.g., sulfate or 5′-methylthioadenosine (MTA) for perturbation) [58].
- Quenching Solution: Liquid nitrogen.
- Extraction Solvent: 10 mM ammonium bicarbonate buffer (pH 9.2) containing 90% acetonitrile [58].
- Instrumentation: High-resolution Liquid Chromatography-Fourier Transform Mass Spectrometry (LC-FTMS) system, e.g., 11T LTQ-FT Ultra [58].
Protocol:
- Cell Culture and Perturbation: Grow model organisms (e.g., Rhodospirillum rubrum, Bacillus subtilis, or microalgae) to mid-log phase (OD~600~ = 0.6-6.0). Apply perturbation (e.g., 1 mM MTA feeding, light/dark treatment) and collect samples at critical time points (e.g., 0, 10, 20 minutes) [58] [59].
- Sample Quenching and Metabolite Extraction:
- Rapidly pellet cells and immediately freeze in liquid nitrogen.
- Resuspend frozen cell pellets in 0.375 mL of ice-cold extraction solvent.
- Vortex vigorously and centrifuge to remove cell debris. Transfer the supernatant for analysis [58].
- LC-FTMS Analysis:
- Perform chromatographic separation using a reversed-phase column.
- Acquire high-resolution mass spectra in negative ionization mode.
- Use quality control samples (pooled from all samples) to monitor instrument performance [58].
Transcriptomics Sample Preparation and qRT-PCR Validation
Objective: To quantify changes in the expression of genes encoding key enzymes in the isoprenoid pathways.
Materials:
- RNA Stabilization: RNase inhibitors or RNA protection kits.
- RNA Extraction Kit: e.g., RNeasy Protect Kit.
- cDNA Synthesis Kit: Reverse transcription kit.
- qPCR Mix: SYBR Green or TaqMan master mix.
- Instrumentation: Real-time PCR thermocycler.
Protocol:
- RNA Extraction:
- cDNA Synthesis:
- Quantify RNA concentration and purity.
- Synthesize cDNA from 1 µg of total RNA using a reverse transcription kit.
- Quantitative Real-Time PCR (qRT-PCR):
- Design primers for key pathway genes (e.g., DXS, DXR, HMGR, TPS) and reference housekeeping genes.
- Prepare reactions in triplicate with SYBR Green master mix, primers, and cDNA template.
- Run qPCR and analyze data using the comparative C~T~ (ΔΔC~T~) method to determine relative gene expression [58] [59].
Data Integration and Computational Analysis
Bioinformatics Pipeline for Data Integration
The logical flow of data from raw outputs to biological insight is depicted below.
Key Quantitative Data from Isoprenoid Pathway Studies
Table 1: Key Metabolite and Gene Expression Changes in Isoprenoid Pathway Studies
| Analyte / Gene | Pathway | Perturbation | Observed Change | Biological Significance |
|---|---|---|---|---|
| MTA | Methionine Salvage / MEP | MTA Feeding (1 mM) | ↑ Accumulation | Precursor influx, perturbs pathway [58] |
| LcDXS3 (Gene) | MEP | Darkness (6h) | 2.76-fold decrease | Key regulatory step suppressed [59] |
| IPP | MVA/MEP Output | Engineered MVA in C. reinhardtii | 7-fold increase (373.75 ± 7.18 µg/g DCW) | Successful heterologous pathway expression [4] |
| LcHMGS1 (Gene) | MVA | Darkness (12h) | 2.63-fold decrease | Cytoplasmic precursor synthesis impaired [59] |
| Non-mevalonate Pathway | MEP | MTA Feeding | Repressed | Reveals unexpected pathway interconnection [58] |
Table 2: Key Enzymes in the Isoprenoid Biosynthetic Pathways
| Enzyme | Abbreviation | Pathway | Function | Diagnostic Utility |
|---|---|---|---|---|
| 1-deoxy-D-xylulose 5-phosphate synthase | DXS | MEP | First committed step; rate-limiting | Key indicator of plastidial pathway flux [10] [4] |
| 1-deoxy-D-xylulose-5-phosphate reductoisomerase | DXR | MEP | Second step; also rate-limiting | Target for perturbation diagnosis [4] |
| 3-hydroxy-3-methylglutaryl-CoA reductase | HMGR | MVA | Key regulatory, irreversible step | Indicator of cytoplasmic pathway activity [10] [4] |
| Terpene Synthase | TPS | Downstream | Converts precursors to final terpenoids | Links precursor flux to end-product synthesis [59] |
| Isopentenyl-diphosphate isomerase | IDI | MVA/MEP | Isomerizes IPP DMAPP | Essential for balance of precursor pools [4] |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents and Tools for Integrated Omics Studies
| Item | Function / Application | Example Specifics |
|---|---|---|
| High-Resolution LC-MS System | Untargeted metabolomics; accurate mass detection. | 11T LTQ-FT Ultra MS; Negative ion mode [58] |
| RNA Protection & Extraction Kit | Preservation and isolation of high-quality RNA for transcriptomics. | RNeasy Protect Kit [58] |
| qRT-PCR Reagents | Validation of gene expression for key pathway genes. | SYBR Green master mix, gene-specific primers [59] |
| Pathway Analysis Software | Data pre-processing, annotation, and enrichment analysis. | XCMS (peak detection), MSEA (enrichment) [58] |
| Model Organisms | Systems biology models for pathway perturbation. | R. rubrum, B. subtilis, C. reinhardtii, L. cubeba [58] [10] [59] |
| Stable Isotope Tracers | (Optional) For flux analysis to track carbon through pathways. | ¹³C-labeled glucose or pyruvate to trace MEP pathway flux |
Isoprenoid Biosynthesis Pathway Diagram
Within the field of isoprenoid biosynthetic pathway engineering, the accumulation of toxic intermediates presents a significant bottleneck to achieving high yields. This Application Note addresses the specific challenge of 3-hydroxy-3-methylglutaryl-CoA (HMG-CoA) toxicity in engineered Escherichia coli strains. The mevalonate (MVA) pathway, a common heterologous pathway introduced into E. coli for isoprenoid production, leads to the formation of HMG-CoA. This intermediate is known to be toxic to the host cell, impairing growth and limiting the overall flux towards target compounds such as terpenoids and their derivatives [60]. The strategic mitigation of this toxicity is therefore a critical consideration within broader metabolic engineering research. This document provides detailed protocols and data for two primary intervention strategies: the implementation of enzyme colocalization to prevent intermediate accumulation and the utilization of an energy-efficient archaeal MVA pathway to reduce metabolic burden.
Strategic Approaches and Key Mechanisms
Two principal metabolic engineering strategies have emerged as effective means to combat HMG-CoA toxicity.
Enzyme Colocalization via Sortase A-Mediated Scaffolding
This approach involves the post-translational formation of a multi-enzyme complex on a scaffold protein, creating a metabolic channeling system. By bringing the enzyme that produces HMG-CoA (MvaS) into close proximity with the enzyme that consumes it (MvaE), the local concentration of the intermediate is minimized, thereby reducing its cytoplasmic diffusion and associated toxicity [60]. This method is particularly advantageous as it redirects metabolic flux without requiring genetic disruption.
Implementation of an Energy-Efficient Archaeal MVA Pathway
An alternative strategy involves pathway selection and optimization. The canonical "eukaryotic" MVA pathway requires more ATP than a recently discovered archaeal MVA pathway. The archaeal pathway utilizes different enzymes, such as phosphomevalonate dehydratase (PMDh) and isopentenyl phosphate kinase (IPK), and consumes two-thirds of the ATP required by the canonical pathway [5]. This energy-saving nature reduces the metabolic burden on the cell and can be exploited to enhance the productivity of isoprenoid compounds like lycopene and β-farnesene in engineered E. coli [5].
The following diagram illustrates the logical workflow for selecting and implementing these strategies.
Experimental Protocols
Protocol 1: SrtA-Mediated Enzyme Assembly on Multimeric Scaffolds
This protocol details the procedure for forming metabolic enzyme complexes on scaffold proteins (Streptavidin, SA, or E. coli CutA, CA) to enhance mevalonate production and reduce HMG-CoA toxicity [60].
Strain and Plasmid Construction
- Step 1: Clone the genes encoding the key enzymes for mevalonate biosynthesis (atoB, mvaE, and mvaS) into expression vectors. The mvaE and mvaS genes should be modified to include a C-terminal LPETG tag for Sortase A recognition.
- Step 2: Clone the genes for the scaffold proteins (SA or CA) with an N-terminal G tag into a separate expression vector. Co-express the srtA gene (encoding Sortase A) in the same vector or a compatible one.
- Step 3: Assemble the final plasmids. The example strains from the research are:
- B/Mlps: Contains pMevlp (expressing AtoB, MvaE-LPETG, MvaS-LPETG) and pZAsrtAs (expressing SrtA and G-SA).
- B/Mlpc: Contains pMevlp and pZAsrtAc (expressing SrtA and G-CA).
Cultivation and Mevalonate Production
- Step 1: Inoculate the engineered E. coli strain (e.g., BW25113) into LB medium with appropriate antibiotics (e.g., 100 μg/mL ampicillin, 50 μg/mL kanamycin). Incubate overnight at 37°C with shaking at 180 rpm.
- Step 2: Sub-culture the pre-culture into M9Y minimal medium (supplemented with antibiotics and 20 g/L glucose) at an initial OD₆₀₀ of approximately 0.05.
- Step 3: Induce protein expression when the OD₆₀₀ reaches 0.5–0.8 by adding IPTG to a final concentration of 0.1 mM.
- Step 4: Continue incubation for 48 hours at 37°C with shaking at 200 rpm.
- Step 5: Analyze mevalonate titer using high-performance liquid chromatography (HPLC) or LC-MS.
Protocol 2: Exploiting the Archaeal MVA Pathway for Isoprenoid Production
This protocol describes the expression of the complete archaeal MVA pathway from Methanosarcina mazei in E. coli for the production of isoprenoids like lycopene and β-farnesene [5].
Pathway Assembly and Strain Engineering
- Step 1: Assemble the upper and lower parts of the archaeal MVA pathway. The pathway genes from M. mazei include MM_1762 (mevalonate kinase, MVK), MM_1525/MM_1524 (subunits of phosphomevalonate dehydratase, PMDh), MM_1526 (anhydromevalonate phosphate decarboxylase, AMPD), MM_1871 (phosphomevalonate synthase, PFS), and MM_1763 (isopentenyl phosphate kinase, IPK).
- Step 2: For lycopene production, co-express the crtE, crtI, and crtB genes on a separate plasmid (e.g., pACYC-IBE).
- Step 3: For β-farnesene production, co-express genes for farnesyl pyrophosphate synthase (FPS) and farnesene synthase (FS).
Cultivation and Product Analysis
- Step 1: Grow the engineered E. coli strain in a suitable rich medium (e.g., TB) or minimal medium with necessary antibiotics.
- Step 2: Induce pathway expression with an appropriate inducer (e.g., L-arabinose for pBAD vectors) when the culture reaches the mid-exponential phase (OD₆₀₀ ~ 2.5–3.0).
- Step 3: For oxygen-sensitive enzymes like PMDh, consider performing cultures under oxygen-limiting or semi-anaerobic conditions to maximize pathway activity [5].
- Step 4: Extract and quantify the target isoprenoid. Lycopene can be measured spectrophotometrically, while β-farnesene can be analyzed using gas chromatography (GC) or GC-MS.
Data Presentation and Analysis
Quantitative Comparison of Strategy Outcomes
The following tables summarize key performance data from the implementation of the described strategies.
Table 1: Performance of SrtA-Mediated Scaffolding for Mevalonate Production [60]
| E. coli Strain | Scaffold Type | Mevalonate Titer (mM) | Relative Improvement vs Control |
|---|---|---|---|
| B/Mn (No Scaffold) | N/A | ~12.5 | 1.0x (Baseline) |
| B/Mlp (SrtA only) | N/A | ~16.5 | ~1.3x |
| B/Mlp_s | Streptavidin (SA) | ~23.5 | ~1.9x |
| B/Mlp_c | CutA (CA) | ~25.0 | ~2.0x |
Table 2: Isoprenoid Production via the Archaeal MVA Pathway in E. coli [5]
| Isoprenoid Product | Cultivation Condition | Productivity (Comparative Level) | Key Pathway Features |
|---|---|---|---|
| Lycopene (Lc) | Semi-anaerobic | Enhanced | Oxygen-sensitive PMDh; functions effectively in low O₂ |
| Lycopene (Lc) | Aerobic | Negligible | PMDh inactivated by oxidation |
| β-Farnesene (Fn) | Defined conditions | Comparable to canonical pathway | Lower ATP consumption (by ~2/3) |
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for Combating HMG-CoA Toxicity
| Reagent / Material | Function / Role in Experiment | Key Details / Considerations |
|---|---|---|
| pMevlp Plasmid | Expresses core mevalonate pathway enzymes (AtoB, MvaE-LPETG, MvaS-LPETG) | Base plasmid for SrtA-mediated scaffolding strategy [60] |
| pZAsrtAs / pZAsrtAc | Expresses Sortase A and scaffold proteins (G-tagged SA or CA) | Enables post-translational assembly of enzyme complexes on scaffolds [60] |
| Archaeal MVA Pathway Genes | Provides an energy-efficient route to IPP/DMAPP | Genes from M. mazei: MM_1762, MM_1525/1524, MM_1526, MM_1871, MM_1763 [5] |
| C-terminal LPETG Tag | Sortase A recognition motif | Fused to MvaE and MvaS for enzymatic ligation to scaffolds [60] |
| N-terminal G Tag | Oligoglycine acceptor for SrtA | Fused to scaffold proteins (SA, CA) for enzyme attachment [60] |
Pathway Diagram
The diagram below illustrates the engineered metabolic pathways for isoprenoid production in E. coli, highlighting the two main strategies for mitigating HMG-CoA toxicity.
Strategies for Reducing Unfavorable Byproducts and Competing Pathways
In the field of isoprenoid biosynthetic pathway engineering, the redirection of cellular resources toward the desired high-value product is a fundamental challenge. Competing endogenous metabolic pathways and the formation of unfavorable byproducts significantly limit the yield and productivity of target isoprenoids in microbial cell factories [10] [61]. This application note details proven metabolic engineering strategies and experimental protocols to overcome these limitations, providing researchers with practical methodologies to enhance flux through engineered isoprenoid pathways. The content is framed within the broader context of optimizing microbial systems for the sustainable production of pharmaceutical, nutraceutical, and industrial isoprenoids.
Core Engineering Strategies
Metabolic engineering approaches to minimize competing pathways focus on redirecting carbon flux toward isoprenoid biosynthesis and preventing substrate depletion by native metabolism [61]. The five primary strategies include: overproduction of enzymes in rate-limited reactions; knockout of genes in competing pathways; creation of new metabolic routes for precursor production; ensuring adequate cofactor supply; and downregulation of reactions leading to undesirable metabolites [10].
Table 1: Strategic Approaches to Minimize Competing Pathways
| Strategy | Engineering Approach | Key Enzymes/Pathways Targeted | Expected Outcome |
|---|---|---|---|
| Precursor Pool Enhancement | Overexpression of rate-limiting enzymes [10] [4] | DXS, DXR, IDI (MEP pathway); HMGR, HMGS, MVK (MVA pathway) [4] | 2.5–7 fold increase in IPP/DMAPP supply [4] |
| Competitive Pathway Knockout | CRISPR-Cas9 gene disruption [10] | Pyruvate dehydrogenase, fatty acid synthesis, endogenous terpene synthases | Redirection of carbon flux from acetyl-CoA and pyruvate to isoprenoids |
| Heterologous Pathway Implementation | Introduction of complete MVA pathway in MEP-only hosts [5] [4] | mvaE, mvaS, mvk, pmk, mvaD from various organisms | Bypass of native regulation; 2.5–3 fold yield improvement [4] |
| Cofactor Balancing | NADPH/ATP regeneration systems [36] | NAD kinase, ATP-generating modules | Improved stoichiometric efficiency in phosphorylation-intensive steps |
| Enzyme Level Optimization | Quantitative tuning of enzyme expression ratios [36] | MVK:PMK:MVD:IDI:ISPS (1:1:1:2:16 optimal ratio) | 5-fold increase in conversion efficiency [36] |
The following diagram illustrates the logical relationship between engineering strategies and their intended effects on the isoprenoid biosynthetic network:
Diagram: Logical flow of engineering strategies to enhance isoprenoid production. Each strategy addresses a specific limitation in the native metabolic network.
Quantitative Performance Data
Engineering efforts to reduce competing pathway effects have demonstrated significant improvements in isoprenoid yields across different microbial platforms. The following table summarizes representative quantitative data from published studies:
Table 2: Quantitative Impact of Pathway Engineering on Isoprenoid Production
| Host Organism | Engineering Strategy | Target Product | Yield Improvement | Reference |
|---|---|---|---|---|
| Escherichia coli | Archaeal MVA pathway implementation | Lycopene, β-farnesene | Comparable to canonical MVA pathway with reduced ATP cost [5] | [5] |
| S. cerevisiae | Alternative precursor utilization (isopentenol) | Linalool & analogs | Established non-canonical building block pathway [57] | [57] |
| C. reinhardtii | Heterologous MVA pathway introduction | IPP/DMAPP precursors | 7-fold increase in IPP (373.75 ± 7.176 μg/g DCW) [4] | [4] |
| In vitro enzyme system | Optimized enzyme ratio (MVK:PMK:MVD:IDI:ISPS) | Isoprene | 5-fold increase (26 mg/L/h) [36] | [36] |
| E. coli | Balanced lower MVA pathway | Isoprene | 430 mg/L/h with 10 balanced enzyme units [36] | [36] |
Detailed Experimental Protocols
Protocol: Competitive Pathway Gene Knockout Using CRISPR-Cas9
Purpose: To disrupt genes in competing metabolic pathways that divert carbon from isoprenoid biosynthesis.
Materials:
- CRISPR-Cas9 plasmid system specific to host organism
- Host-specific transformation reagents
- Selection antibiotics appropriate for CRISPR system
- Synthetic guide RNA (sgRNA) targeting competing pathway genes
- DNA oligonucleotides for homology-directed repair (HDR) template (if using)
Procedure:
- sgRNA Design: Design sgRNAs to target key enzymes in competing pathways such as pyruvate dehydrogenase complex (aceEF), fatty acid synthesis (fabH), or endogenous terpene synthases [10].
- Plasmid Construction: Clone validated sgRNAs into appropriate CRISPR-Cas9 expression vector with selection marker.
- Transformation: Introduce CRISPR plasmid into host strain using optimized transformation method (electroporation for bacteria, lithium acetate for yeast).
- Selection and Screening: Plate transformed cells on selective media and incubate until colonies appear.
- Verification: Screen colonies by colony PCR and sequence verification to confirm gene disruptions.
- Plasmid Curing: Remove CRISPR plasmid through counter-selection or serial passage in non-selective media.
- Phenotypic Validation: Assess impact on growth characteristics and measure precursor pool changes.
Protocol: Enzyme Ratio Optimization for In Vitro Isoprenoid Production
Purpose: To quantitatively balance enzyme levels in multi-enzyme systems for maximized flux to target isoprenoids.
Materials:
- Purified enzymes for targeted pathway (e.g., MVK, PMK, MVD, IDI, ISPS)
- Substrates (mevalonate, ATP)
- Reaction buffer (50 mM Tris-HCl, pH 7.5, 10 mM MgCl₂)
- ATP regeneration system (if applicable)
- GC-MS system for isoprene detection
Procedure:
- Initial Screening: Set up 2 mL reaction systems with constant levels of four enzymes (0.5 μM each) while varying the fifth enzyme from 0.02-5 μM to identify bottleneck enzymes [36].
- Rate Determination: Measure initial production rates at each enzyme level to identify concentration where reaction rate plateaus.
- Ratio Optimization: Based on bottleneck identification, test different molar ratios of all five enzymes centered on optimal individual concentrations.
- Validation at Scale: Scale up optimized system to 50 mL volume with proportional enzyme units.
- Continuous Monitoring: Sample reactions at multiple time points (e.g., 0, 2, 4, 8, 24, 40 h) to determine production kinetics.
- Data Analysis: Calculate production rates and compare to unbalanced control system (0.5 μM each enzyme).
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Isoprenoid Pathway Engineering
| Reagent/Category | Specific Examples | Function/Application |
|---|---|---|
| CRISPR Systems | Cas9 expression vectors, sgRNA scaffolds | Targeted gene knockout of competing pathways [10] |
| Pathway Enzymes | MVK, PMK, MVD, IDI, ISPS, DXS, DXR, HMGR | Rate-limiting enzyme overexpression; in vitro pathway reconstruction [36] [4] |
| Analytical Standards | MVA, MVAP, MVAPP, IPP, DMAPP, GPP, FPP | LC-MS quantification of pathway intermediates [62] |
| Extraction Solvents | Acetonitrile:50mM ammonium formate (7:3, pH 9.5) | Efficient extraction of mevalonate pathway metabolites [62] |
| Heterologous Pathways | Archaeal MVA pathway genes; mvaE, mvaS from E. faecalis | Bypass native regulatory mechanisms; enhance precursor supply [5] [4] |
| Promoter Systems | Inducible (PGAL1, Para); constitutive strong promoters | Fine-tuned control of enzyme expression levels [57] |
Pathway Engineering Workflow
The following diagram outlines a comprehensive experimental workflow for implementing strategies to reduce competing pathways in isoprenoid engineering:
Diagram: Comprehensive workflow for engineering reduced competition in isoprenoid pathways. The iterative process enables continuous optimization until performance targets are met.
Concluding Remarks
Effective reduction of unfavorable byproducts and competing pathways requires a systematic approach combining multiple metabolic engineering strategies. The protocols and data presented herein provide a foundation for researchers to design and implement pathway engineering projects aimed at enhancing isoprenoid production. As the field advances, integration of machine learning for pathway prediction and enzyme design will further accelerate the development of efficient microbial cell factories for isoprenoid biosynthesis [63] [33].
Platforms in Perspective: A Comparative Analysis of Engineering Hosts
Within metabolic engineering, the selection of a microbial host is a pivotal decision that directly impacts the titers, rates, and yields (TRY) of bioproducts. Isoprenoids, a vast class of natural compounds with applications ranging from pharmaceuticals to biofuels, present a particular challenge and opportunity for biosynthetic production. Efficiently channeling carbon through complex metabolic pathways to a desired isoprenoid requires a host that is not only genetically tractable but also physiologically compatible with the target molecule and its precursors. This application note provides a structured comparison of common and emerging microbial hosts used in isoprenoid pathway engineering, detailing quantitative performance metrics and standard protocols for benchmarking production.
Host Organism Comparison and Selection
Microbial hosts vary significantly in their inherent physiological properties, genetic toolkits, and metabolic capabilities. The table below summarizes the key characteristics of several organisms employed in isoprenoid biosynthesis.
Table 1: Comparison of Microbial Hosts for Isoprenoid Production
| Host Organism | Key Advantages | Key Disadvantages | Exemplary Isoprenoid / Metric (Value) | Primary Isoprenoid Pathway |
|---|---|---|---|---|
| Escherichia coli | Fast growth, high genetic tractability, well-understood physiology [64] | Lack of native isoprenoid pathway, cytotoxicity from pathway intermediates [64] | Amorpha-4,11-diene (precursor to artemisinin) [64] | Heterologous Mevalonate (MVA) Pathway [64] |
| Saccharomyces cerevisiae | Native MVA pathway, efficiently expresses eukaryotic proteins, robust industrial host [65] | Limited precursor supply, competitive sterol biosynthesis, sensitivity to stressful conditions [65] | General terpenoids and isoprenoids [65] | Native MVA Pathway [65] |
| Pichia pastoris | High cell-density fermentation, efficient protein secretion, minimal glycosylation, strong methanol-induced promoters [66] | Less developed genetic tools compared to S. cerevisiae, requirement for methanol induction [66] [67] | Menaquinone-4 (MK-4) (0.24 mg/g DCW) [67] | Native MVA Pathway [67] |
| Microalgae (e.g., C. reinhardtii, P. tricornutum) | CO₂ fixation as carbon source, photosynthetic, compartmentalization (e.g., chloroplasts) [10] | Slow growth, challenging genetic manipulation, limited precursor availability [10] | Carotenoids, phytosterols [10] | MEP, MVA, or both (species-dependent) [10] |
Detailed Experimental Protocols for Host Evaluation
To ensure reproducible and comparable results when benchmarking different microbial hosts, standardized protocols for cultivation and analysis are critical.
Protocol: Cultivation and Induction of RecombinantPichia pastorisfor MK-4 Production
This protocol is adapted from the study that achieved a menaquinone-4 (MK-4) titer of 0.24 mg/g DCW in P. pastoris [67].
Strains and Vectors:
- Host Strain: P. pastoris GS115 (a histidine-deficient strain with Mut+ phenotype) [67].
- Expression Vector: pGAPZα, enabling constitutive expression under the GAP promoter, or pPICZα, for methanol-inducible expression under the AOX1 promoter [67].
- Gene of Interest: HsUBIAD1, encoding the human prenyltransferase responsible for converting menadione (VK₃) to MK-4 [67].
Culture Media and Conditions:
- Seed Culture: Inoculate a single colony into 25 mL of BMGY medium (1% yeast extract, 2% peptone, 1.34% YNB, 1% glycerol, 4×10⁻⁵% biotin, 0.1 M potassium phosphate pH 6.0) in a 250 mL flask. Incubate at 30°C with shaking at 250 rpm for approximately 24 hours until the OD₆₀₀ reaches a suitable density [67].
- For Constitutive Expression (pGAPZα): Dilute the seed culture to an OD₆₀₀ of ~0.05 in fresh BMGY medium and continue cultivation for ~24 hours [67].
- For Inducible Expression (pPICZα):
- Harvest cells from the seed culture by centrifugation (5,000 × g, 10 min, room temperature).
- Resuspend the cell pellet in BMMY medium (same as BMGY but with 0.3% methanol instead of glycerol) to an OD₆₀₀ of 1.0.
- Maintain induction by adding methanol to a final concentration of 0.3% every 24 hours [67].
- Optimal Conditions for HsUBIAD1: The highest yield was observed at pH 7.0 and 24°C after 36 hours of incubation [67].
Analytical Method:
- MK-4 Quantification: Extract isoprenoids from cell pellets using appropriate organic solvents (e.g., hexane). Analyze extracts via High-Performance Liquid Chromatography (HPLC) or LC-MS/MS. Quantify MK-4 against a standard curve of purified MK-4 and normalize to Dry Cell Weight (DCW) [67].
Protocol: Investigating and Counteracting Intermediate Toxicity inE. coli
This protocol is based on the study that identified HMG-CoA accumulation as a cytotoxic event in an engineered E. coli strain and implemented a supplementation strategy to mitigate it [64].
Strain and Pathway Engineering:
Culture Conditions:
- Grow engineered strains in defined C medium supplemented with 3.4% glycerol and necessary antibiotics [64].
- Induce pathway expression by adding 1.33 mM (0.02%) L-arabinose when the culture OD₆₀₀ reaches ~0.2-0.3 [64].
- To counteract HMG-CoA toxicity, supplement the medium with 100 μg/mL palmitic acid (16:0) or oleic acid (cis-Δ9-18:1) along with 400 μg/mL Brij detergent to facilitate solubilization [64].
Analysis of Toxicity and Rescue:
- Growth Kinetics: Monitor OD₆₀₀ over time to quantify the growth defect upon induction and the recovery with fatty acid supplementation [64].
- Transcriptomics: Use DNA microarray analysis to profile global gene expression changes. The study identified the inhibition of the type II fatty acid biosynthesis (FAB) pathway as the primary mode of toxicity [64].
- Targeted Metabolite Profiling: Employ LC-MS/MS to quantify intracellular levels of pathway intermediates, specifically HMG-CoA, confirming its accumulation [64].
Pathway Visualization and Experimental Workflow
The biosynthesis of all isoprenoids originates from the universal five-carbon precursors, isopentenyl pyrophosphate (IPP) and dimethylallyl pyrophosphate (DMAPP). Microbial hosts utilize one of two major pathways to produce these building blocks.
Diagram 1: Core Isoprenoid Biosynthesis Pathways in Microbial Hosts. The MVA pathway is native to most eukaryotes like yeast, while the MEP pathway is found in bacteria and the plastids of microalgae. Engineered E. coli often employs a heterologous MVA pathway.
A standard workflow for benchmarking production across hosts involves constructing the pathway, analyzing performance, and iteratively optimizing the system.
Diagram 2: Workflow for Benchmarking Isoprenoid Production. This iterative process integrates systems biology data to diagnose and resolve production bottlenecks, such as intermediate toxicity or precursor limitation.
The Scientist's Toolkit: Key Reagents and Solutions
Table 2: Essential Research Reagents for Isoprenoid Pathway Engineering
| Reagent / Material | Function / Application | Example & Notes |
|---|---|---|
| pBAD Series Vectors | Inducible (arabinose) expression of pathway genes in E. coli [64] | pBAD33MevT used to express the mevalonate pathway; allows regulated gene expression to manage toxicity [64]. |
| pGAPZα / pPICZα Vectors | Constitutive (GAP) or inducible (AOX1) expression in P. pastoris [66] [67] | pGAPZα avoids methanol use; pPICZα enables strong, methanol-induced expression [67]. |
| Fatty Acids (Palmitic/Oleic) | Counteract toxicity from pathway intermediates in E. coli [64] | 100 μg/mL with Brij detergent rescues growth inhibition from HMG-CoA by replenishing fatty acid pools [64]. |
| Methanol (HPLC Grade) | Inducer for the AOX1 promoter in P. pastoris [66] | Typically used at 0.5-2.0% concentration; requires strict process control for safety [66]. |
| HsUBIAD1 Enzyme | Key prenyltransferase for menaquinone-4 (MK-4) synthesis [67] | Catalyzes the prenylation of menadione (VK₃) with geranylgeranyl diphosphate (GGPP) to form MK-4 [67]. |
| CRISPR/Cas9 Systems | Targeted genome editing for pathway optimization and gene knockout [66] [10] | Enables precise engineering in yeasts and microalgae to enhance precursor flux or delete competing pathways [66] [10]. |
Isoprenoids represent one of the most diverse classes of natural compounds, with applications spanning pharmaceuticals, nutraceuticals, biofuels, and fragrances [10] [68]. Traditional microbial production hosts, such as Escherichia coli and Saccharomyces cerevisiae, have been extensively engineered for isoprenoid biosynthesis but face limitations including high feedstock costs and precursor flux constraints [30] [34]. Microalgae, photosynthetic microorganisms, have emerged as promising alternative chassis due to their inherent capabilities for solar energy utilization and metabolic compartmentalization [10] [69]. This Application Note details the comparative advantages of microalgae and provides foundational protocols for leveraging these organisms in isoprenoid pathway engineering research.
Comparative Analysis: Microalgae vs. Conventional Hosts
The following table summarizes the key operational and metabolic differences between microalgae and conventional hosts that are critical for isoprenoid production.
Table 1: Key Feature Comparison between Conventional Hosts and Microalgae for Isoprenoid Production
| Feature | Conventional Hosts (E. coli, S. cerevisiae) | Microalgae (e.g., C. reinhardtii, P. tricornutum) |
|---|---|---|
| Carbon Source | Organic feedstocks (e.g., glucose, glycerol); costly and contributes >90% of production expenses [30]. | Inorganic CO₂; provides a sustainable, carbon-neutral substrate [10] [70]. |
| Energy Source | Chemical energy from substrate catabolism. | Light energy via photosynthesis; minimizes need for organic carbon [10] [70]. |
| Native Isoprenoid Pathways | Typically one: MVA in yeast, MEP in E. coli [71]. | Often native MEP pathway; some diatoms possess both MEP and MVA pathways [10]. |
| Compartmentalization | Limited in prokaryotes; present in yeast but less plastid-specific. | Chloroplast enables targeted engineering and isolation of metabolic pathways/harmful intermediates [10]. |
| Cultivation Costs & Sustainability | High-cost sugar feedstocks; competition with food sources. | Can be cultivated on non-arable land using wastewater; lower freshwater footprint [10]. |
| Genetic Toolkits | Highly advanced and extensive synthetic biology tools available. | Tools are developing rapidly; CRISPR/Cas systems and omics resources are increasingly available [72] [10]. |
Advantages of the Photosynthetic Platform
Microalgae convert light energy and CO₂ into chemical energy and carbon skeletons, directly fueling the metabolic network that produces isoprenoids.
Table 2: Quantitative Advantages of Microalgal Photosynthesis for Bioproduction
| Parameter | Terrestrial Plants | Microalgae | Significance for Production |
|---|---|---|---|
| Solar Conversion Efficiency | 1-2% [73] | Up to 4.4% [73] | Higher efficiency translates to faster growth rates and greater biomass yields. |
| Theoretical Max Biomass Productivity | Lower (seasonal dependence) [74] | ~80 g/m²/day or 280 ton/ha/year [70] | Enables higher annual yields of intracellular bioproducts like isoprenoids. |
| Land Requirement | High (arable land) | Negligible (can use marginal lands or photobioreactors) [70] | Does not compete with food production; suitable for scalable biorefinery concepts. |
The diagram below illustrates how photosynthesis integrates with and fuels the isoprenoid biosynthetic pathway in microalgae.
Advantages of Metabolic Compartmentalization
The chloroplast, the site of photosynthesis, is also a self-contained metabolic factory. It houses the entire MEP pathway for IPP and DMAPP synthesis in green microalgae [10]. This allows for:
- Targeted Engineering: Metabolic pathways can be engineered directly within the chloroplast by using plastid-specific promoters and transformation techniques, minimizing metabolic crosstalk with the cytosol [10].
- Precursor Proximity: The MEP pathway operates in the chloroplast, utilizing the G3P and pyruvate generated directly by the Calvin cycle and glycolysis within the same organelle, enhancing metabolic flux [10] [70].
- Sequestration of Toxic Intermediates: Compartmentalization allows harmful pathway intermediates or final products to be isolated, reducing cellular toxicity and feedback inhibition [10].
Detailed Experimental Protocols
Protocol 1: Chloroplastic Transformation of C. reinhardtii for Pathway Engineering
This protocol describes the delivery of genetic constructs to the chloroplast for overexpression of MEP pathway genes.
I. Materials
- Chlamydomonas reinhardtii strain (e.g., CC-503 cw92 mt+)
- Spectinomycin stock solution (50 mg/mL in H₂O)
- Tris-Acetate-Phosphate (TAP) media, solid and liquid
- Gene gun (e.g., Biolistic PDS-1000/He system) or glass bead transformation setup
- Gold or tungsten microparticles (0.6 µm)
- Chloroplast transformation vector (e.g., pASapI) containing:
- Plastid 16S rRNA promoter (Prrn) and psbA 3' UTR for transgene expression.
- Spectinomycin resistance gene (aadA) as a selectable marker.
- Gene of interest (e.g., DXR or IspS)
II. Methods
- Algal Cultivation: Grow C. reinhardtii in liquid TAP media under continuous light (50 µE m⁻² s⁻¹) with shaking until mid-log phase (cell density ~2-5 x 10⁶ cells/mL).
- Particle Preparation: a. Weigh 60 mg of gold microparticles into a 1.5 mL tube. b. Add 1 µg of purified plasmid DNA, 100 µL of 2.5 M CaCl₂, and 40 µL of 0.1 M spermidine. Vortex continuously for 10 minutes. c. Let particles settle, remove supernatant, and wash with 1 mL of 100% ethanol. d. Resuspend particles in 100 µL of 100% ethanol.
- Transformation: a. Concentrate 2 x 10⁸ algal cells by centrifugation and resuspend in 300 µL of TAP medium. Spread as a thin layer in the center of a TAP agar plate. b. For gene gun transformation, coat the macrocarrier with the DNA-particle suspension and perform the shot according to the manufacturer's instructions (e.g., 1,100 psi rupture disc, 6 cm target distance). c. For glass bead transformation, mix concentrated cells with DNA and acid-washed glass beads and vortex vigorously for 30 seconds.
- Selection and Screening: a. After transformation, incubate plates under light for 24 hours in the dark to allow for gene expression. b. Transfer cells to TAP plates containing 100 µg/mL spectinomycin. c. Incubate under light for 2-3 weeks until green colonies appear. d. Pick resistant colonies and sub-culture repeatedly on selective media to achieve homoplasmy (complete replacement of the chloroplast genome).
III. Analysis
- Confirm integration of the transgene into the chloroplast genome via PCR and Southern blotting.
- Measure transcript levels of the introduced gene using RT-qPCR.
- Quantify isoprenoid product yield using GC-MS or HPLC (see Protocol 3).
Protocol 2: Cultivation and Induction of Isoprenoid Production in P. tricornutum
This protocol leverages the unique dual MEP/MVA pathway in the diatom Phaeodactylum tricornutum for high-level isoprenoid production.
I. Materials
- Phaeodactylum tricornutum strain (e.g., UTEX 646)
- f/2 medium (artificial seawater supplemented with nutrients)
- Bioreactor or environmental shaker with controlled light and temperature
| Reagent | Function |
|---|---|
| Silicate Solution | Essential nutrient for diatom cell wall formation [70]. |
| Vitamin B12, Biotin, Thiamine | Essential micronutrients in f/2 medium for robust growth. |
| Nitrogen (NaNO₃) & Phosphorus (NaH₂PO₄) | Macronutrients; depletion can be used to trigger lipid and isoprenoid accumulation [10]. |
| High-Intensity LED Lighting | Provides controllable, uniform light energy for photosynthesis. |
II. Methods
- Inoculum Preparation: Inoculate P. tricornutum from a fresh agar plate into 50 mL of f/2 medium. Grow under continuous light (100 µE m⁻² s⁻¹) at 22°C with shaking until the culture reaches the late-log phase.
- Scale-Up Cultivation: Transfer the inoculum to a 2L photobioreactor containing 1.5L of f/2 medium. Maintain temperature at 22°C, pH at 7.5, and provide continuous light and aeration (0.5 vvm, air with 2% CO₂).
- Stress Induction: Once the culture reaches a high cell density (stationary phase, ~5 x 10⁷ cells/mL), induce isoprenoid production by one of two methods: a. Nutrient Stress: Harvest cells by gentle centrifugation, wash, and resuspend in f/2 medium lacking nitrogen (N-) or phosphorus (P-). b. High-Light Stress: Increase light intensity to 500-1000 µE m⁻² s⁻¹.
- Harvest: Continue cultivation under stress conditions for 3-5 days, then harvest cells by centrifugation for product analysis.
III. Analysis
- Monitor growth by measuring optical density at 750 nm (OD₇₅₀) and cell count.
- Extract and quantify target isoprenoids (e.g., fucoxanthin) as described in Protocol 3.
Protocol 3: Analytical Quantification of Isoprenoids via HPLC
A standard method for the extraction and quantification of carotenoids and other isoprenoids from microalgal biomass.
I. Materials
- Lyophilized microalgal biomass
- Methanol, Acetone, Ethyl Acetate (HPLC grade)
- Diethyl Ether
- Solid Phase Extraction (SPE) cartridges (e.g., C18)
- High-Performance Liquid Chromatography (HPLC) system with a photodiode array (PDA) detector
- C18 reverse-phase column (e.g., 250 mm x 4.6 mm, 5 µm)
II. Methods
- Extraction: a. Weigh 10 mg of freeze-dried algal powder into a 2 mL tube. b. Add 1.5 mL of extraction solvent (e.g., methanol:ethyl acetate, 1:1 v/v) and 0.5 g of glass beads (0.5 mm). c. Homogenize using a bead beater for 5 minutes at 30 Hz. d. Centrifuge at 13,000 x g for 10 minutes. Transfer the supernatant to a new tube. e. Repeat the extraction on the pellet until it becomes colorless. Pool all supernatants. f. Evaporate the combined extract to dryness under a stream of nitrogen gas.
- Sample Preparation: Redissolve the dry extract in 200 µL of methanol and filter through a 0.22 µm syringe filter before HPLC analysis.
- HPLC Analysis: a. Column: C18 reverse-phase. b. Mobile Phase: A: Water with 0.1% Formic Acid; B: Acetonitrile with 0.1% Formic Acid. c. Gradient: 0 min: 70% B; 0-20 min: 70% B → 100% B; 20-25 min: 100% B; 25-26 min: 100% B → 70% B. d. Flow Rate: 1.0 mL/min. e. Detection: PDA detector, acquire spectra from 200-600 nm. Quantify using external standards based on absorbance at specific wavelengths (e.g., 450 nm for carotenoids).
III. Analysis
- Identify compounds by comparing retention times and UV-Vis spectra to authentic standards.
- Quantify concentration using a calibration curve constructed from standard solutions.
The Scientist's Toolkit: Key Research Reagents
Essential materials for engineering isoprenoid biosynthesis in microalgae.
Table 3: Essential Research Reagents for Microalgal Isoprenoid Engineering
| Reagent / Tool Category | Specific Example(s) | Function / Application |
|---|---|---|
| Model Microalgae Species | Chlamydomonas reinhardtii [72] [70] | A genetically tractable model organism for green algae. |
| Phaeodactylum tricornutum [72] [10] | A diatom model with available genetic tools and both MEP/MVA pathways. | |
| Genetic Engineering Tools | CRISPR/Cas9 systems [72] [10] | Enables precise genome editing for gene knockouts or knock-ins. |
| Chloroplast-specific promoters (e.g., Prrn, psbA) [10] | Drives high-level transgene expression in the chloroplast. | |
| Omics & Computational Resources | Genome-scale Metabolic Models (GEMs) [73] | In silico prediction of metabolic fluxes and identification of engineering targets. |
| AlgaePath database [72] | Aids in metabolic pathway prediction and analysis. | |
| Cultivation & Induction | f/2 Medium [70] | Standard culture medium for marine microalgae like diatoms. |
| Nitrogen-depleted Medium [10] | A stress condition used to trigger lipid and isoprenoid accumulation. |
Microalgae present a compelling and sustainable alternative to conventional microbial hosts for isoprenoid production. Their ability to utilize sunlight and CO₂, coupled with the metabolic advantages of chloroplast compartmentalization, addresses key economic and technical hurdles in pathway engineering. The application notes and detailed protocols provided here offer a roadmap for researchers to harness the unique capabilities of microalgae, paving the way for the development of efficient, solar-driven cell factories for the production of high-value isoprenoids.
Isoprenoids represent one of the most diverse classes of natural products with applications spanning pharmaceuticals, nutraceuticals, flavors, fragrances, and biofuels [30] [10] [75]. The commercial viability of microbial isoprenoid production faces significant challenges, primarily driven by substrate costs which can account for over 90% of total production expenses [30]. Conventional microbial chassis such as Escherichia coli and Saccharomyces cerevisiae typically utilize sugar substrates like glucose and glycerol, which provide high metabolic flux through the mevalonate (MVA) or methylerythritol phosphate (MEP) pathways but contribute to prohibitive production costs at industrial scale [30] [76].
Recent research has challenged conventional isoprenoid production paradigms by exploring atypical carbon substrates including C1 compounds (methane, methanol, carbon dioxide, formate), waste streams (cellulosic biomass, glycerol), and other non-sugar carbon sources [30] [76] [77]. This application note provides a comprehensive economic and sustainability analysis comparing traditional sugar substrates with emerging atypical alternatives, framed within the context of isoprenoid biosynthetic pathway engineering. We present quantitative comparisons, detailed experimental protocols, and visual workflow representations to facilitate adoption of these sustainable approaches within research and industrial settings.
Carbon Substrate Economics in Isoprenoid Biosynthesis
Quantitative Analysis of Substrate Options
Table 1: Economic and Metabolic Comparison of Carbon Substrates for Isoprenoid Production
| Carbon Substrate | Theoretical Carbon Yield | Reported Isoprenoid Yield | Cost Relative to Glucose | Sustainability Advantages |
|---|---|---|---|---|
| Glucose | Base | High titers reported [30] | 1.0 (reference) | Renewable but competes with food sources |
| Soybean Oil | Higher than sugars [76] | 0.72–0.76 g PHA/g substrate [76] | Not specified | Waste streams available |
| Glycerol | Similar to sugars | 25-28x higher conversion in E. coli vs. native producers [76] | Lower than glucose | Industrial byproduct |
| C1 Substrates (CO₂, methanol) | Variable | Under characterization [30] | Potentially much lower | Carbon capture/utilization |
| Lignocellulosic Biomass | Similar to glucose | 53% cost reduction potential [30] | ~47% of glucose cost [30] | Non-food biomass, abundant waste streams |
The economic advantage of atypical substrates extends beyond simple feedstock costs. Substrates like fatty acids and vegetable oils demonstrate superior carbon economy due to more direct metabolic routing to key isoprenoid precursors. For instance, soybean oil achieves yields of 0.72–0.76 g PHA per gram of substrate in Ralstonia eutropha cultures, significantly higher than yields typically obtained from sugar substrates [76]. This enhanced yield stems from the more efficient conversion of oil-derived acetyl-CoA to isoprenoid precursors compared to the oxidative glycolysis pathway required for sugar conversion, which inherently loses carbon as CO₂ [76].
Metabolic and Sustainability Considerations
Beyond direct economic factors, substrate selection profoundly influences environmental impact and process sustainability. The use of waste streams like glycerol (a biodiesel byproduct), discarded vegetable oil, or pretreated cellulosic biomass reduces reliance on food-competing substrates and contributes to circular bioeconomy models [30] [76]. Additionally, C1 substrates like methanol and CO₂ offer the potential for carbon-negative or carbon-neutral production systems when coupled with renewable energy [30].
Metabolic challenges differ significantly between substrate classes. Sugar metabolism often faces precursor limitations and feedback inhibition issues [30] [12], while C1 metabolism requires specialized pathways and enzymes not present in conventional chassis [30]. Organisms capable of metabolizing atypical substrates, such as methylotrophs and phototrophic microbes, often possess inherent tolerance to inhibitory compounds present in waste streams, providing an additional advantage for industrial processes [30].
Experimental Protocols for Atypical Substrate Utilization
Isopentenol Utilization Pathway (IUP) Protocol
The Isopentenol Utilization Pathway represents a paradigm-shifting approach that decouples isoprenoid production from central carbon metabolism [12] [57]. This two-step pathway utilizes externally provided isopentenol isomers as substrates instead of sugar-derived catabolites.
Table 2: Key Research Reagent Solutions for IUP Implementation
| Reagent/Component | Function | Examples/Specifications |
|---|---|---|
| Isopentenol Isomers | Pathway substrate | Isoprenol or prenol (0.6-1 mM initial concentration) |
| First-step Kinases | Phosphorylation of isopentenol | ScCK (S. cerevisiae), AtFKI (A. thaliana) |
| Second-step Kinases | Diphosphate formation | AtIPK (A. thaliana), MtIPK (M. thermautotrophicus) |
| IDI Enzyme | IPP/DMAPP balancing | Isopentenyl-pyrophosphate delta isomerase |
| Expression System | Heterologous pathway expression | Inducible (pTET) or constitutive (Ppro4) promoters |
Protocol: IUP Implementation in E. coli
Strain Engineering
- For proof-of-concept: Create MEP-knockout strain (e.g., KO1 with ispG deletion) using CRISPR-Cas9 system [12]
- Transform with IUP plasmid: pSEVA228-pro4IUPi (constitutive) or pTET-IUPi (inducible) expressing ScCK, AtIPK, and IDI
- Include appropriate antibiotic resistance markers for selection
Culture Conditions
- Use minimal media (e.g., M9) supplemented with 0.6 mM isoprenol or prenol
- Maintain temperature at 37°C with appropriate agitation (220 rpm)
- For inducible systems, add anhydrotetracycline (aTC) at mid-log phase
Analytical Validation
- Monitor growth kinetics with spectrophotometry (OD600)
- Quantify isoprenoid production (e.g., lycopene via UV/Vis spectroscopy)
- Confirm IPP/DMAPP production via LC-MS [78]
Pathway Optimization
- Fine-tune expression levels using RBS engineering
- Balance IPP:DMAPP ratio by modulating IDI expression
- Supplement with ATP precursors if needed
Figure 1: Isopentenol Utilization Pathway for Isoprenoid Biosynthesis. This two-step pathway converts isopentenol isomers to IPP and DMAPP using sequential phosphorylation, bypassing native metabolic routes [12] [57].
Methylotrophic Platform Protocol
Methylotrophic organisms utilize C1 compounds such as methanol and methane, offering exceptional carbon economy and sustainability profiles.
Protocol: Engineering Methylotrophic Isoprenoid Production
Host Selection and Engineering
- Select methylotrophic host (e.g., Methylorubrum extorquens, Methanosarcina sp.)
- Engineer MVA or MEP pathway amplification using strong native promoters
- Introduce heterologous terpene synthases for target isoprenoids
Culture Conditions
- Use minimal media with methanol (0.5-1%) as sole carbon source
- Maintain temperature according to host requirements (typically 30-37°C)
- Control methanol feeding rate to prevent toxicity
- For methane-utilizing systems, optimize gas-liquid transfer
Process Monitoring
- Track methanol consumption via HPLC or enzymatic assays
- Quantify isoprenoid production using GC-MS or HPLC
- Monitor key pathway intermediates (IPP, DMAPP) [78]
Strain Optimization
- Apply adaptive laboratory evolution for improved methanol tolerance
- Use CRISPR interference for downregulation of competing pathways
- Engineer cofactor balancing for improved NADPH regeneration
Phototrophic Production Protocol
Phototrophic microorganisms like cyanobacteria and microalgae utilize CO₂ as their carbon source, providing the most direct route to carbon-neutral isoprenoid production.
Protocol: CO₂-based Isoprenoid Production in Microalgae
Strain Selection and Transformation
- Select microalgal host with genetic tractability (e.g., Chlamydomonas reinhardtii, Phaeodactylum tricornutum)
- Engineer MEP pathway (present in plastids) via nuclear or chloroplast transformation
- Express heterologous terpene synthases targeted to plastids
Culture Conditions
- Use BG-11 or similar minimal medium with CO₂ enrichment (2-5%)
- Provide continuous illumination (50-100 μmol photons/m²/s)
- Maintain temperature at 25°C with continuous mixing
Metabolic Engineering Strategies
- Overexpress rate-limiting enzymes (DXS, IDI) [10]
- Knock out competing pathways (fatty acid synthesis)
- Engineer cofactor supply (ATP, NADPH) via light-harvesting modifications
Product Recovery
- Harvest cells during stationary phase
- Implement extraction protocols (solvent, supercritical CO₂)
- Purify target isoprenoids using chromatographic methods
Integrated Analysis and Future Perspectives
The transition from sugar-based to atypical carbon substrates for isoprenoid production represents both a technical challenge and strategic opportunity for the biotechnology industry. Our analysis demonstrates that substrates such as methanol, CO₂, and waste streams offer compelling economic advantages with potential cost reductions exceeding 50% compared to glucose-based systems [30]. Furthermore, the sustainability benefits of these alternatives align with growing emphasis on circular bioeconomy principles and carbon-neutral manufacturing.
Figure 2: Integrated Workflow for Isoprenoid Production from Diverse Carbon Sources. The diagram illustrates the metabolic connection between substrate choice, pathway engineering, and final product synthesis [30] [76] [77].
Technical implementation requires careful consideration of both host organism selection and pathway engineering strategy. While conventional hosts like E. coli and S. cerevisiae benefit from extensive engineering toolkits, non-model organisms often possess innate metabolic capabilities for atypical substrate utilization that can be leveraged through modern genetic engineering approaches [30]. The development of synthetic pathways like IUP further expands the possibilities by decoupling isoprenoid production from native central metabolism [12] [57].
Future research should focus on enhancing carbon conversion efficiency, expanding the range of utilizable waste streams, and developing integrated biorefinery approaches that co-produce isoprenoids with other valuable bioproducts. Advances in enzyme engineering, particularly for C1 assimilation pathways and promiscuous terpene synthases, will further accelerate adoption of these sustainable platforms [57]. Additionally, life cycle assessment studies are needed to fully quantify the environmental benefits of atypical substrate utilization across the complete production chain.
The protocols and analyses presented herein provide a foundation for researchers to implement and advance these approaches, contributing to more economically viable and environmentally sustainable isoprenoid production systems.
Application Notes
Within isoprenoid biosynthetic pathway engineering, a central challenge is selecting an appropriate chassis organism, navigating the core trade-off between genetic tractability and native resilience. Isoprenoids, the largest class of natural products, are essential for numerous pharmaceuticals, nutraceuticals, and industrial applications [10] [13]. Their structural complexity makes chemical synthesis challenging, driving research towards microbial and plant-based bioproduction [79] [10].
Genetic tractability refers to the ease with which an organism's genetic code can be modified, a characteristic of established platforms like Escherichia coli and Saccharomyces cerevisiae. In contrast, native resilience describes the inherent capacity of an organism to withstand metabolic and environmental stresses, often due to complex, evolved physiological and biochemical networks [79] [80]. This trade-off directly impacts the yield, stability, and economic viability of engineered isoprenoid pathways.
Table 1: Comparative Analysis of Isoprenoid Production Chassis
| Chassis Characteristic | High Tractability Platforms (e.g., E. coli, S. cerevisiae) | Native Resilience Platforms (e.g., Microalgae, Plants) |
|---|---|---|
| Typical Engineering Cycle (Design-Build-Test-Learn) | Rapid; well-established tools [79] | Slower; transformation and regeneration can be bottlenecks [79] [81] |
| Heterologous Enzyme Expression | May struggle with complex plant-derived enzymes [79] | Naturally accommodates intricate metabolic networks [79] |
| Tolerance to Compound Toxicity | Often low; can impair cell growth and yield [79] | High; natural compartmentalization (e.g., vacuoles, plastids) mitigates toxicity [79] [10] |
| Precursor Availability (e.g., IPP/DMAPP) | Requires extensive pathway engineering (MVA or MEP) [10] | Often native MEP and/or MVA pathways present; can be optimized [10] |
| Environmental Resilience | Low; requires sterile, controlled conditions [10] | High; can be cultivated on non-arable land with wastewater [10] |
| Carbon Source | Organic feedstocks (costly) [10] | Atmospheric CO₂ (sustainable) [10] |
Key Experimental Approaches for Investigation
Researchers employ several strategies to evaluate and overcome the trade-offs in chassis selection:
- Integrated Omics and Genome Editing for Functional Validation: Combining genomics, transcriptomics, and metabolomics identifies key genes and regulatory nodes in resilient species. CRISPR/Cas systems are then used for precise knockout or activation of these targets. For example, CRISPR/Cas9 was used to edit glutamate decarboxylase genes in tomato, leading to a 7- to 15-fold increase in GABA accumulation [79].
- Transient Expression in Hybrid Systems: Platforms like Nicotiana benthamiana balance tractability and plant-native resilience. Its rapid biomass and efficient transient transformation allow quick testing of multi-gene pathways, such as the reconstruction of diosmin (involving 5-6 enzymes) or QS-7 saponin (19 genes) biosynthesis [79].
- Metabolic Engineering to Overcome Precursor Limitation: In both tractable and resilient hosts, a common goal is to enhance the carbon flux toward isoprenoid precursors (IPP and DMAPP). This involves overexpressing rate-limited enzymes, knocking out competitive pathways, and ensuring adequate cofactor supply [10].
- Leveraging Nanotechnology to Bolster Resilience: Nanoparticles (NPs) like zinc oxide (ZnO) and magnesium oxide (MgO) can be applied to plants to enhance innate abiotic stress tolerance. They function by scavenging reactive oxygen species (ROS) and improving nutrient uptake, thereby stabilizing yields under drought or salinity stress, which protects the engineered metabolic pathways [82] [81].
Protocols
Protocol 1: Rapid Pathway Prototyping Using aNicotiana benthamianaTransient Expression System
This protocol is designed to quickly test the functionality of a reconstructed isoprenoid pathway in a plant chassis, leveraging its relative tractability and native cellular machinery.
1. Principle Agrobacterium tumefaciens is used to deliver T-DNA constructs containing the genes of interest into leaf cells. The plant's transcriptional machinery then drives transient expression, allowing for rapid assessment of enzyme activity and metabolite production within a cellular environment that possesses native resilience features like compartmentalization [79].
2. Reagents and Equipment
- Nicotiana benthamiana plants (4-5 weeks old)
- Agrobacterium tumefaciens strain (e.g., GV3101) harboring the expression vector(s) with your isoprenoid pathway genes
- YEP broth and agar plates with appropriate antibiotics
- Infiltration buffer: 10 mM MES, 10 mM MgCl₂, 100 µM acetosyringone (pH 5.6)
- 1 mL needleless syringe
- Liquid nitrogen for sample freezing
- Analytical equipment (LC-MS, GC-MS) for metabolite detection
3. Step-by-Step Procedure Day 1: Culture Initiation Inoculate a single colony of each Agrobacterium strain into 5 mL of YEP broth with antibiotics. Incubate at 28°C with shaking (200 rpm) for 24 hours.
Day 2: Culture Preparation and Infiltration 1. Sub-culture the bacteria into fresh YEP medium (with antibiotics) to an OD₆₀₀ of 0.1. Grow to an OD₆₀₀ of 0.6-1.0. 2. Pellet cells by centrifugation (5000 × g, 10 min). Resuspend the pellet in infiltration buffer to a final OD₆₀₀ of 0.4-0.6. 3. For multi-gene pathways, combine equal volumes of different bacterial suspensions. 4. Incubate the bacterial suspension at room temperature for 2-4 hours. 5. Using a needleless syringe, press the tip against the abaxial side of a N. benthamiana leaf and gently inject the suspension. Infiltrate multiple leaves/plants per construct.
Day 4-7: Harvest and Analysis 1. Harvest the infiltrated leaf tissue, flash-freeze in liquid nitrogen, and store at -80°C. 2. Lyophilize or homogenize the tissue under liquid nitrogen. 3. Extract metabolites using a suitable solvent (e.g., methanol or ethyl acetate). 4. Analyze the extract for target isoprenoids using LC-MS/GC-MS.
Protocol 2: Enhancing Microalgal Isoprenoid Yield via CRISPR-Mediated Gene Knock-In
This protocol outlines a strategy to improve the native resilience and biosynthetic capacity of microalgae, a chassis that requires more advanced genetic tool development.
1. Principle Microalgae like Chlamydomonas reinhardtii possess native MEP pathways for isoprenoid biosynthesis and are highly resilient, growing photoautotrophically on CO₂. This protocol uses CRISPR/Cas9 to integrate a heterologous gene (e.g., a rate-limiting enzyme from the mevalonate pathway) into a genomic "safe harbor" or a specific metabolic gene locus, thereby optimizing carbon flux without disrupting essential functions [10].
2. Reagents and Equipment
- Chlamydomonas reinhardtii culture
- CRISPR/Cas9 plasmid system for the selected microalga
- Donor DNA template for homologous recombination
- Cell wall-deficient strain or enzyme for cell wall digestion
- Electroporator or gene gun
- Tris-Acetate-Phosphate (TAP) medium, solid and liquid
- Selection antibiotics (e.g., paromomycin, hygromycin)
- PCR reagents for genotyping
3. Step-by-Step Procedure Step 1: Target Selection and Construct Design 1. Identify the target locus (e.g., a gene in the MEP pathway or a safe harbor). 2. Design and clone a sgRNA targeting this locus into the CRISPR/Cas9 vector. 3. Design a donor DNA template containing your gene of interest, flanked by homology arms (~500-800 bp) to the target site.
Step 2: Transformation 1. Grow C. reinhardtii to mid-log phase in TAP medium. 2. If using a walled strain, digest cell walls to generate protoplasts. 3. Co-deliver the CRISPR/Cas9 plasmid and the donor DNA via electroporation or particle bombardment. 4. Allow recovery in liquid TAP without selection for 24 hours.
Step 3: Selection and Screening 1. Plate cells on TAP agar containing the appropriate antibiotic. 2. After 1-2 weeks, pick resistant colonies and transfer to fresh plates. 3. Perform PCR genotyping on the colonies to confirm precise gene integration at the target locus.
Step 4: Phenotypic Validation 1. Grow confirmed positive lines in photobioreactors under standard and stress conditions (e.g., high light, salinity). 2. Quantify growth metrics and extract isoprenoids for yield analysis. 3. Compare yields to wild-type and empty vector controls to assess improvement.
Visualizations
Diagram 1: Trade-offs in Chassis Selection for Isoprenoid Engineering
Diagram 2: Isoprenoid Biosynthesis and Engineering Nodes
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Isoprenoid Pathway Engineering
| Reagent / Tool | Function / Application | Example in Context |
|---|---|---|
| CRISPR/Cas9 System | Targeted gene knockout, knock-in, or activation in resilient chassis. | Editing SlGAD genes in tomato to increase GABA [79]; engineering microalgal MEP pathway genes [10]. |
| Agrobacterium tumefaciens (GV3101) | Delivery of T-DNA for stable or transient plant transformation. | Transient expression of multi-gene pathways for diosmin or saponin biosynthesis in N. benthamiana [79]. |
| Metabolomics Standards | Quantitative analysis of pathway intermediates and final products. | LC-MS/MS or GC-MS analysis of IPP/DMAPP, carotenoids, or reconstructed compounds like diosmin [79] [10]. |
| Nanoparticles (ZnO, MgO) | Priming agents to enhance abiotic stress tolerance in plants. | Foliar application to improve plant health and stabilize yields under drought/salinity, protecting engineered pathways [82] [81]. |
| Design-Build-Test-Learn (DBTL) Software | Computational framework for iterative cycle design and modeling. | Predicting metabolic flux, optimizing enzyme combinations, and learning from failed experiments to refine strategies [79]. |
The escalating climate crisis, driven by increasing concentrations of greenhouse gases, necessitates a fundamental transformation in industrial biomanufacturing [83] [84]. The transition from traditional, sugar-based feedstocks to sustainable one-carbon (C1) substrates represents a paradigm shift toward a circular bioeconomy. This application note examines two classes of microbial hosts with significant potential for sustainable production: methylotrophs, which can utilize C1 compounds like methanol, and oleaginous yeasts, renowned for their high lipid accumulation [83] [85] [86]. Within the specific context of isoprenoid biosynthetic pathway engineering, we explore how these platforms can be harnessed and optimized for the carbon-neutral production of high-value terpenoids. Isoprenoids, a vast family of over 55,000 natural products including pharmaceuticals, fragrances, and potential biofuels, are traditionally biosynthesized from hemiterpene building blocks via metabolically expensive and tightly regulated native pathways [53]. Engineering future-proof platforms involves decoupling production from agricultural feedstocks, enhancing carbon conversion efficiency, and creating orthogonal biosynthetic systems to overcome native metabolic limitations [53] [84]. This document provides a detailed technical overview of current engineering paradigms, quantitative performance data, and standardized protocols for advancing research in this field.
Methylotrophic Platforms: Chassis for C1 Valorization
Methylotrophic organisms are capable of utilizing reduced C1 compounds such as methanol as a sole carbon and energy source. Their utility lies in the sustainable nature of methanol, which can be produced from syngas, methane, or captured CO₂ [83] [86].
- Native Methylotrophs: Organisms like Methylorubrum extorquens, Bacillus methanolicus, and the yeast Pichia pastoris possess innate C1 assimilation pathways. For instance, B. methanolicus has been reported to produce up to 59 g/L of L-glutamate from methanol [86]. However, they often suffer from a limited genetic toolbox and insufficient knowledge of their regulatory networks.
- Synthetic Methylotrophs: To overcome the limitations of native hosts, significant efforts have been directed toward engineering synthetic methylotrophy in industrially proven chassis like Escherichia coli and Saccharomyces cerevisiae [83] [87]. Recent breakthroughs have yielded an engineered E. coli strain capable of growth on methanol with a doubling time of 4.3 hours, a performance comparable to many natural methylotrophs [87].
The core metabolism of methylotrophs involves methanol oxidation to formaldehyde, its detoxification/dissimilation to CO₂ for energy, and its assimilation into biomass. The three main assimilation pathways are the Ribulose Monophosphate (RuMP) cycle, the Xylulose Monophosphate (XuMP) cycle, and the Serine cycle [86]. The RuMP cycle, used by many bacteria, is considered particularly efficient for biomass yield, as it assimilates carbon at the oxidation level of formaldehyde and requires only one ATP per pyruvate generated [86].
Oleaginous Yeasts: Powerhouses for Lipid and Isoprenoid Synthesis
Oleaginous yeasts are defined by their ability to accumulate high levels of neutral lipids (over 20-30% of their dry cell weight), making them ideal platforms for lipid-derived chemicals and fuels [85] [88]. Their natural robustness, ability to grow on diverse and inexpensive feedstocks, and good growth at low pH (which prevents bacterial contamination) are significant advantages for industrial bioprocessing [85].
- Conventional Hosts: Yarrowia lipolytica is the most well-studied oleaginous yeast, with a extensive suite of genetic tools available. It has been successfully engineered for the production of organic acids, polyunsaturated fatty acids (PUFAs), and alkanes [85].
- Emerging Hosts: Yeasts from the genera Rhodosporidium (e.g., R. toruloides), Trichosporon, and Lipomyces are gaining attention due to their ability to achieve even higher lipid accumulation (over 60%) and their adaptability to a wide range of substrates, including lignocellulosic hydrolysates [85] [88]. The genus Rhodosporidium is also a natural producer of carotenoids [85].
The inherent high flux of acetyl-CoA in oleaginous yeasts, the central precursor for both fatty acids and mevalonate (MVA) pathway-derived isoprenoids, positions them as superior chassis for terpenoid biosynthesis [85]. Recent advances in systems biology and genetic tool development are now unlocking the potential of these non-conventional yeasts [88].
Table 1: Comparative Analysis of Promising Microbial Platforms for Isoprenoid Biosynthesis
| Platform Organism | Key Feature | Production Example (Non-Isoprenoid) | Genetic Toolbox | Relevance to Isoprenoid Engineering |
|---|---|---|---|---|
| Native Methylotrophs (M. extorquens) | Utilizes methanol natively | N/A | Limited | Potential for direct C1-to-terpenoid conversion; pathway understanding is key [86]. |
| Synthetic Methylotrophs (E. coli) | Engineered methanol utilization | 1 g/L Itaconic acid [87] | Extensive | Chassis for methanol-based terpenoid production using established engineering tools [87]. |
| Oleaginous Yeast (Y. lipolytica) | High lipid accumulation; robust | ~90 g/L Citric acid [85] | Advanced | Excellent host due to high acetyl-CoA supply; engineered for terpenoids [85] [88]. |
| Oleaginous Yeast (R. toruloides) | Very high lipid accumulation; inhibitor tolerant | Carotenoids [85] | Developing | Emerging host; high natural precursor pool for lipids and isoprenoids [85]. |
Engineering and Application in Isoprenoid Biosynthesis
Engineering Strategies for Enhanced Precursor Supply
A primary bottleneck in microbial isoprenoid production is the supply of the universal five-carbon building blocks, isopentenyl diphosphate (IPP) and dimethylallyl diphosphate (DMAPP). Native pathways like the Mevalonate (MVA) and Methylerythritol Phosphate (MEP) pathways are complex, highly regulated, and energetically costly [53]. Two key engineering strategies have emerged to address this.
1. Decoupling Hemiterpene Biosynthesis via an Artificial Pathway: An innovative approach involves designing an artificial, orthogonal hemiterpene pathway. The proposed Alcohol-Dependent Hemiterpene (ADH) pathway starts with membrane-permeable alcohols like isopentenol (ISO) and dimethylallyl alcohol (DMAA). These are converted to IPP and DMAPP via a two-step phosphorylation catalyzed by a promiscuous phosphatase (Kinase-1, e.g., PhoN) and an isopentenyl phosphate kinase (IPK, Kinase-2) [53]. This pathway is completely decoupled from native metabolism, requires only two ATPs (compared to more in native pathways), and allows for the potential use of non-natural alcohol precursors to create novel isoprenoid analogues [53]. This system has been successfully coupled to lycopene and prenylated tryptophan biosynthetic systems in E. coli [53].
2. Rewiring Central Carbon Metabolism: In both methylotrophic and oleaginous hosts, enhancing the flux of central carbon precursors (acetyl-CoA, glyceraldehyde-3-phosphate, pyruvate) into the MVA or MEP pathways is critical. This can be achieved by:
- Overexpression of Bottleneck Enzymes: In Y. lipolytica, overexpressing enzymes like acetyl-CoA carboxylase (ACC) and diacylglycerol acyltransferase (DGA2) enhances flux towards acetyl-CoA-derived products [85].
- Pathway Amplification: Heterologous expression of the entire MVA pathway in the cyanobacterium Synechocystis resulted in a 2.5-fold increase in photosynthetic isoprene production by overcoming the flux limitations of the native MEP pathway [89].
- Carbon Conservation: In methylotrophic E. coli, mutations in the dissimilatory RuMP cycle gene gnd (6-phosphogluconate dehydrogenase) are thought to reduce carbon loss as CO₂, thereby increasing the carbon yield available for product synthesis, including from nodes like acetyl-CoA [87].
Production of Isoprenoids and Other Value-Added Compounds
Engineering these platforms has led to successful demonstrations of chemical production from C1 and non-conventional feedstocks.
Table 2: Exemplary Products from Engineered Methylotrophic and Oleaginous Platforms
| Product Category | Specific Product | Host Organism | Feedstock | Key Engineering Strategy | Reference |
|---|---|---|---|---|---|
| Isoprenoids | Isoprene | Synechocystis sp. PCC 6803 | CO₂ & H₂O (Photo-synthetic) | Heterologous expression of MVA pathway + isoprene synthase | [89] |
| Isoprenoids | α-Farnesene | Yarrowia lipolytica | Glucose/Sucrose | Metabolic pathway engineering | [90] |
| Isoprenoids | β-Farnesene | Pichia pastoris | Methanol | Engineered synthetic methylotrophy | [84] |
| Organic Acids | Lactic Acid | Synthetic Methylotrophic E. coli | Methanol | Production from pyruvate node | [87] |
| Polymers | Polyhydroxy-butyrate (PHB) | Synthetic Methylotrophic E. coli | Methanol | Production from acetyl-CoA node | [87] |
| Aromatics | p-Aminobenzoic Acid (PABA) | Synthetic Methylotrophic E. coli | Methanol | Production from chorismate pathway | [87] |
| Lipids | Fatty Acid Ethyl Esters (FAEE) | Yarrowia lipolytica | Glycerol | Engineering of FA-derived products | [85] |
Detailed Experimental Protocols
Protocol 1: Coupling the Artificial ADH Pathway to Lycopene Production inE. coli
This protocol outlines the steps to implement the orthogonal Alcohol-Dependent Hemiterpene (ADH) pathway to produce lycopene, using a lycopene reporter system to quantify success [53].
Principle: Heterologously expressed kinases (PhoN and IPK) convert supplemented alcohols (DMAA/ISO) to DMAPP/IPP, which are used by the endogenous prenyltransferase machinery and heterologous lycopene pathway enzymes to produce the red pigment lycopene.
Workflow:
Materials:
- Bacterial Strains: E. coli BL21(DE3) or similar.
- Plasmids:
- pCDFDuet-GGPP: Encodes geranylgeranyl diphosphate synthase (CrtE or IspA Y80D).
- pACYCDuet-Lyc: Encodes the lycopene biosynthesis genes (CrtB, CrtI).
- pETDuet-IPK: Co-expresses Thermoplasma acidophilum Isopentenyl Phosphate Kinase (IPK) and Shigella flexneri PhoN phosphatase.
- Reagents: Dimethylallyl alcohol (DMAA), Isopentenol (ISO), Fosmidomycin (Fs), Isopropyl β-d-1-thiogalactopyranoside (IPTG), Lycopene standard.
- Media: M9 minimal media supplemented with appropriate antibiotics (Spectinomycin, Chloramphenicol, Ampicillin).
Procedure:
- Strain Construction: Co-transform the three plasmids (pCDFDuet-GGPP, pACYCDuet-Lyc, pETDuet-IPK) into competent E. coli cells. Select transformants on LB agar plates with Spectinomycin (50 µg/mL), Chloramphenicol (34 µg/mL), and Ampicillin (100 µg/mL).
- Culture and Induction: Inoculate a single colony into M9 media with antibiotics and grow overnight at 37°C. Dilute the culture to an OD600 of ~0.1 in fresh media. Add Fosmidomycin (0.5 µM) to inhibit the native DXP pathway. Induce gene expression with IPTG (0.1-0.5 mM) when OD600 reaches ~0.6. Simultaneously, supplement the culture with DMAA and ISO (each at a final concentration of 2.5 mM).
- Incubation: Incubate the induced culture at 30°C with shaking for 24-48 hours to allow for pigment production.
- Lycopene Extraction: Harvest cells by centrifugation. Resuspend the cell pellet in 1 mL of acetone and incubate at 55°C for 15 minutes to extract lycopene. Centrifuge at high speed to remove cell debris.
- Quantification: Transfer the acetone extract to a new tube. Analyze the lycopene content by HPLC or measure the absorbance at 450-470 nm. Compare against a standard curve of purified lycopene for quantitative data.
Protocol 2: Engineering a Synthetic MethylotrophicE. colifor Bioproduction
This protocol describes the general methodology for utilizing an evolved synthetic methylotrophic E. coli strain for chemical production from methanol [87].
Principle: An engineered E. coli strain (e.g., MEcoliref2) equipped with the RuMP cycle (via genes for methanol dehydrogenase, Hps, and Phi) and adaptive laboratory evolution (ALE) gains the ability to use methanol as a carbon source. Production pathways are then inserted at key metabolic nodes.
Workflow:
Materials:
- Bacterial Strain: Synthetic methylotrophic E. coli MEcoliref2 (or similar) [87].
- Bioreactor: A fully controlled fed-batch bioreactor with pH, dissolved oxygen (DO), and temperature control.
- Media: Defined minimal media. Methanol is used as the sole or primary carbon source.
- Genetic Tools: Plasmids or chromosomal integration systems for introducing heterologous production pathways (e.g., for itaconic acid, PHA, lactic acid).
Procedure:
- Strain Engineering: Introduce the biosynthetic pathway for the target chemical (e.g., itaconic acid, PHB, lactic acid) into the methylotrophic E. coli chassis. This can be achieved via plasmid-based expression or chromosomal integration. Verify gene expression and functionality.
- Fed-Batch Fermentation:
- Inoculate a seed culture of the engineered strain and grow to mid-log phase.
- Transfer the seed culture to the bioreactor containing the defined minimal media.
- Initiate a fed-batch process with a continuous or pulsed feed of methanol to maintain a low, non-toxic concentration (e.g., 300-500 mM initial, allowing consumption and controlled feeding) while preventing excessive evaporation.
- Strictly control process parameters: Temperature = 37°C, pH = 7.0, DO > 30%.
- Monitoring and Harvest: Regularly monitor optical density (OD600) and methanol concentration. The evolved strain MEcoliref2 can achieve high cell densities (OD600 ~100). Harvest the culture when growth plateaus or when product titer is maximized.
- Product Analysis:
- Lactic Acid/PABA: Analyze by HPLC.
- PHB: Quantify by Gas Chromatography (GC) after methanolysis of cell pellets.
- Itaconic Acid: Quantify by HPLC or LC-MS.
The Scientist's Toolkit: Key Research Reagents and Materials
Table 3: Essential Research Reagents for Engineering Methylotrophic and Oleaginous Platforms
| Reagent / Material | Function / Application | Example & Notes |
|---|---|---|
| Methanol Dehydrogenase (Mdh) | Oxidizes methanol to formaldehyde in bacterial methylotrophs. | Bacillus Mdh (NAD+-dependent) or M. extorquens Mdh (PQQ-dependent). Kinetic optimization (e.g., Mdh(H165N, F279I) lowers KM) is crucial [87] [86]. |
| Isopentenyl Phosphate Kinase (IPK) | Second kinase in the artificial ADH pathway; phosphorylates IP to IPP. | From Thermoplasma acidophilum. Allows orthogonal generation of isoprenoid precursors [53]. |
| Promiscuous Phosphatase (PhoN) | First kinase in the artificial ADH pathway; phosphorylates DMAA/ISO to monophosphates. | Class-A non-specific acid phosphatase from Shigella flexneri [53]. |
| Hps/Phi Enzymes | Core of the RuMP cycle for formaldehyde assimilation. | 3-hexulose-6-phosphate synthase (Hps) and 6-phospho-3-hexuloisomerase (Phi). Expression tuning is critical for formaldehyde detoxification [87]. |
| Strong & Inducible Promoters | Drives high-level expression of heterologous genes. | P. pastoris AOX1 (methanol-inducible), Y. lipolytica hybrid promoters (e.g., UAS1B8-TEF), synthetic constitutive promoters [83] [85]. |
| CRISPR-Cas9 System | Enables precise genome editing (gene knock-outs, knock-ins). | Validated systems for Y. lipolytica and R. toruloides for targeted genetic modifications [88]. |
| Fosmidomycin | Inhibits the native DXP pathway for isoprenoid biosynthesis. | Used to suppress native metabolism and isolate the function of engineered pathways [53]. |
Conclusion
The strategic engineering of isoprenoid biosynthetic pathways has matured from a proof-of-concept endeavor to a powerful approach for the sustainable manufacturing of high-value compounds. Success hinges on a holistic strategy that integrates foundational pathway knowledge with advanced metabolic engineering, systematic troubleshooting, and careful host selection. Future directions point toward the increased use of non-model organisms with innate resilience, the expansion of substrate ranges to include cost-effective and sustainable C1 feedstocks, and the application of multi-omics data for intelligent, model-guided strain optimization. For biomedical and clinical research, these advances promise to secure robust and economical supply chains for essential isoprenoid-derived pharmaceuticals, from antimalarials to anticancer agents, while opening new avenues for drug discovery and development.
References
- 1. Two distinct pathways for essential metabolic precursors ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3365244/]
- 2. Towards efficient terpenoid biosynthesis: manipulating IPP ... [https://bioresourcesbioprocessing.springeropen.com/articles/10.1186/s40643-019-0242-z]
- 3. A redundant isoprenoid biosynthetic pathway supports ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12345219/]
- 4. Engineering a complete mevalonate pathway in ... [https://www.sciencedirect.com/science/article/abs/pii/S2211926425000967]
- 5. Enhanced isoprenoid production in Escherichia coli cells ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12547664/]
- 6. The methylerythritol phosphate pathway as an oxidative ... [https://www.nature.com/articles/s41467-024-49483-8]
- 7. Divergent contribution of the MVA and MEP pathways to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10212524/]
- 8. Harnessing the MVA pathway to produce high-value ... [https://portal.nifa.usda.gov/web/crisprojectpages/1032180-harnessing-the-mva-pathway-to-produce-high-value-terpenoids-in-tomato-fruits.html]
- 9. Reconstituted ferredoxin–MEP pathway of Apicomplexa in ... [https://www.sciencedirect.com/science/article/pii/S0021925825025785]
- 10. Progress and prospects in metabolic engineering approaches ... [https://biotechnologyforbiofuels.biomedcentral.com/articles/10.1186/s13068-025-02665-y]
- 11. Analysing intracellular isoprenoid metabolites in diverse ... [https://www.sciencedirect.com/science/article/abs/pii/S0076687922001094]
- 12. Two-step pathway for isoprenoid synthesis - PubMed Central [https://pmc.ncbi.nlm.nih.gov/articles/PMC6329939/]
- 13. The mevalonate pathway of isoprenoid biosynthesis supports ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12265665/]
- 14. The isoprenoid biosynthesis enzyme HDS participates in ... [https://pubmed.ncbi.nlm.nih.gov/41025320/]
- 15. Protocol: analytical methods for visualizing the indolic ... [https://plantmethods.biomedcentral.com/articles/10.1186/s13007-021-00763-0]
- 16. Protocol for analyzing the biosynthesis and degradation of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7890039/]
- 17. Methylerythritol Phosphate Pathway of Isoprenoid ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5031371/]
- 18. The eukaryotic MEP-pathway genes are evolutionarily ... [https://bmcgenomics.biomedcentral.com/articles/10.1186/s12864-021-07448-x]
- 19. Discovery of a metabolic alternative to the classical ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3857490/]
- 20. Advances in the Biosynthesis of Plant Terpenoids [https://pmc.ncbi.nlm.nih.gov/articles/PMC12114759/]
- 21. Progress and prospects in metabolic engineering approaches ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12175399/]
- 22. [ Engineering in Escherichia coli for amorphadiene...] MEP pathway [https://pubmed.ncbi.nlm.nih.gov/24818480/]
- 23. Combination of Entner-Doudoroff Pathway with MEP ... | PLOS One [https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0083290]
- 24. Toward improved terpenoids biosynthesis: strategies to ... [https://bioresourcesbioprocessing.springeropen.com/articles/10.1186/s40643-022-00493-8]
- 25. Biosynthesis of Edible Terpenoids: Hosts and Applications [https://pmc.ncbi.nlm.nih.gov/articles/PMC11854313/]
- 26. Systematic biotechnological production of isoprenoid analogs ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11873216/]
- 27. Biotechnological production of terpenoids using cell factories [https://www.frontiersin.org/journals/pharmacology/articles/10.3389/fphar.2025.1621406/full]
- 28. Optimization of the mevalonate-based isoprenoid biosynthetic ... [https://pubmed.ncbi.nlm.nih.gov/18775787/]
- 29. Review Catalytic mechanism and engineering of aromatic ... [https://www.sciencedirect.com/science/article/abs/pii/S014181302504766X]
- 30. Diversifying Isoprenoid Platforms via Atypical Carbon ... [https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2021.791089/full]
- 31. Microbial Platform for Terpenoid Production: Escherichia ... [https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2018.02460/full]
- 32. Distributing a metabolic pathway among a microbial ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4867547/]
- 33. Synthetic biology and metabolic engineering paving the way ... [https://pubs.rsc.org/en/content/articlehtml/2025/ya/d5ya00118h]
- 34. Metabolic engineering strategies to overcome precursor ... [https://www.sciencedirect.com/science/article/abs/pii/S0958166920300938]
- 35. Metabolic Engineering of Terpenoid Biosynthesis in ... [https://www.mdpi.com/1467-3045/47/9/723]
- 36. Enzymatic process optimization for the in vitro production of ... [https://microbialcellfactories.biomedcentral.com/articles/10.1186/s12934-016-0622-4]
- 37. Chloroplastic metabolic engineering coupled with ... [https://www.nature.com/articles/s41467-019-12879-y]
- 38. Shortcut for isoprenoid biosynthesis | Nature Catalysis [https://www.nature.com/articles/s41929-019-0240-8]
- 39. Isoprenoids Biosynthesis - an overview [https://www.sciencedirect.com/topics/biochemistry-genetics-and-molecular-biology/isoprenoids-biosynthesis]
- 40. Applications of CRISPR/Cas9 in the Synthesis ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7905030/]
- 41. Microbial Genome Editing with CRISPR–Cas9 [https://www.mdpi.com/2311-5637/11/7/410]
- 42. Beyond Cutting: CRISPR-Driven Synthetic Biology Toolkit for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12347677/]
- 43. Enhancing isoprenol production by systematically tuning ... [https://www.frontiersin.org/journals/bioengineering-and-biotechnology/articles/10.3389/fbioe.2023.1296132/full]
- 44. Metabolic engineering and synthetic biology for isoprenoid ... [https://pubmed.ncbi.nlm.nih.gov/33394155/]
- 45. Pathway engineering for functional isoprenoids [https://www.sciencedirect.com/science/article/abs/pii/S0958166911000164]
- 46. Metabolic engineering of Escherichia coli using CRISPR– ... [https://www.sciencedirect.com/science/article/abs/pii/S1096717615000750]
- 47. Biotechnological approaches for artemisinin production in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5871647/]
- 48. Biotechnological Approaches for Production of Artemisinin ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9103073/]
- 49. Safeguarding artemisinin: integrating bioproduction ... [https://malariajournal.biomedcentral.com/articles/10.1186/s12936-025-05648-4]
- 50. Stable Production of the Antimalarial Drug Artemisinin in ... [https://www.frontiersin.org/journals/bioengineering-and-biotechnology/articles/10.3389/fbioe.2017.00047/full]
- 51. Microbial cell factories for bio-based isoprenoid production ... [https://www.sciencedirect.com/science/article/abs/pii/S2452310023000598]
- 52. Metabolic engineering for the high-yield production of ... [https://www.nature.com/articles/srep11128]
- 53. An artificial pathway for isoprenoid biosynthesis decoupled ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6556385/]
- 54. Metabolic Engineering of the Isopentenol Utilization ... [https://www.mdpi.com/1660-3397/20/9/577]
- 55. Engineering acetyl-CoA supply and ERG9 repression to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8788843/]
- 56. Metabolic Flux Analysis of Plastidic Isoprenoid Biosynthesis in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4012595/]
- 57. Systematic biotechnological production of isoprenoid ... [https://www.nature.com/articles/s41467-025-57494-2]
- 58. Integration of untargeted metabolomics with ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4334135/]
- 59. A Comprehensive Analysis of Transcriptomics and ... [https://www.mdpi.com/1422-0067/26/7/2992]
- 60. Sortase A-Mediated Enzyme Assembly on Multimeric ... [https://www.sciepublish.com/article/pii/334]
- 61. Metabolic engineering strategies to overcome precursor ... [https://pubmed.ncbi.nlm.nih.gov/32853882/]
- 62. Optimization of extraction conditions for LC-ToF-MS ... [https://www.sciencedirect.com/science/article/abs/pii/S003991402200978X]
- 63. Comprehensive evaluation of the capacities of microbial ... [https://www.nature.com/articles/s41467-025-58227-1]
- 64. Application of Functional Genomics to Pathway Optimization for Increased Isoprenoid Production - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC2394933/]
- 65. The Model System Saccharomyces cerevisiae Versus ... [https://www.mdpi.com/2075-1729/10/11/299]
- 66. Advances in Metabolic Engineering of Pichia pastoris ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10608127/]
- 67. Construction of a novel MK-4 biosynthetic pathway in Pichia ... [https://microbialcellfactories.biomedcentral.com/articles/10.1186/s12934-019-1215-9]
- 68. Protein engineering strategies for microbial production of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7322351/]
- 69. Genetic Engineering of Microalgae for Secondary Metabolite ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8984019/]
- 70. Potential and Challenges of Improving Photosynthesis in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7020468/]
- 71. Isoprenoid-Based Biofuels: Homologous Expression and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5038050/]
- 72. Bioengineering of Microalgae - PubMed Central - NIH [https://pmc.ncbi.nlm.nih.gov/articles/PMC7494788/]
- 73. Charting the state of GEMs in microalgae [https://www.frontiersin.org/journals/plant-science/articles/10.3389/fpls.2025.1614397/full]
- 74. Metabolic engineering for the production of isoprene and ... [https://link.springer.com/article/10.1007/s00253-018-9200-5]
- 75. Creating plant molecular factories for industrial and ... [https://www.sciencedirect.com/science/article/abs/pii/S0958166917300964]
- 76. Systematic Engineering for Improved Carbon Economy in ... [https://www.mdpi.com/1996-1944/11/8/1271]
- 77. Alternative Carbon Sources for Isoprene Emission - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC6354897/]
- 78. Functional analysis of isoprenoid precursors biosynthesis ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6704079/]
- 79. Plant synthetic biology: from knowledge to biomolecules - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC12602438/]
- 80. A review of plants strategies to resist biotic and abiotic ... [https://www.sciencedirect.com/science/article/abs/pii/S0048969723044571]
- 81. Strategies for Enhancing Resilience in Horticultural Crops ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12415420/]
- 82. Crop management to enhance plant resilience to abiotic ... [https://www.frontiersin.org/journals/plant-science/articles/10.3389/fpls.2025.1626624/full]
- 83. Synthetic methylotrophic yeasts for the sustainable fuel and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9587593/]
- 84. Recent advances in the biosynthesis of natural products ... [https://www.sciencedirect.com/science/article/abs/pii/S1004954125001478]
- 85. Metabolic Engineering of Oleaginous Yeasts for Production of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5682390/]
- 86. Developing methylotrophic microbial platforms for a ... [https://www.frontiersin.org/journals/bioengineering-and-biotechnology/articles/10.3389/fbioe.2022.1050740/full]
- 87. A synthetic methylotrophic Escherichia coli as a chassis for ... [https://www.nature.com/articles/s41929-024-01137-0]
- 88. Recent developments of oleaginous yeasts toward ... [https://www.osti.gov/pages/biblio/2567022]
- 89. Heterologous Expression of the Mevalonic Acid Pathway in ... [https://www.sciencedirect.com/science/article/pii/S167420521460856X]
- 90. Metabolic Engineering of Oleaginous Yeasts for Production ... [https://www.osti.gov/pages/biblio/1510464]